A single-cell atlas of pig gastrulation as a resource for comparative embryology
- PMID: 38890321
- PMCID: PMC11189408
- DOI: 10.1038/s41467-024-49407-6
A single-cell atlas of pig gastrulation as a resource for comparative embryology
Abstract
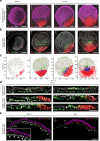
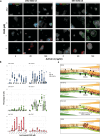
Cell-fate decisions during mammalian gastrulation are poorly understood outside of rodent embryos. The embryonic disc of pig embryos mirrors humans, making them a useful proxy for studying gastrulation. Here we present a single-cell transcriptomic atlas of pig gastrulation, revealing cell-fate emergence dynamics, as well as conserved and divergent gene programs governing early porcine, primate, and murine development. We highlight heterochronicity in extraembryonic cell-types, despite the broad conservation of cell-type-specific transcriptional programs. We apply these findings in combination with functional investigations, to outline conserved spatial, molecular, and temporal events during definitive endoderm specification. We find early FOXA2 + /TBXT- embryonic disc cells directly form definitive endoderm, contrasting later-emerging FOXA2/TBXT+ node/notochord progenitors. Unlike mesoderm, none of these progenitors undergo epithelial-to-mesenchymal transition. Endoderm/Node fate hinges on balanced WNT and hypoblast-derived NODAL, which is extinguished upon endodermal differentiation. These findings emphasise the interplay between temporal and topological signalling in fate determination during gastrulation.
© 2024. Crown.
Conflict of interest statement
The authors declare no competing interests.
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

