Phospholipid Signaling in Crop Plants: A Field to Explore
- PMID: 38891340
- PMCID: PMC11174929
- DOI: 10.3390/plants13111532
Phospholipid Signaling in Crop Plants: A Field to Explore
Abstract
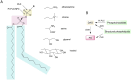
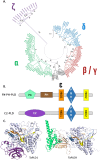
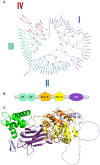
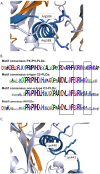
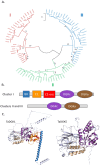
In plant models such as Arabidopsis thaliana, phosphatidic acid (PA), a key molecule of lipid signaling, was shown not only to be involved in stress responses, but also in plant development and nutrition. In this article, we highlight lipid signaling existing in crop species. Based on open access databases, we update the list of sequences encoding phospholipases D, phosphoinositide-dependent phospholipases C, and diacylglycerol-kinases, enzymes that lead to the production of PA. We show that structural features of these enzymes from model plants are conserved in equivalent proteins from selected crop species. We then present an in-depth discussion of the structural characteristics of these proteins before focusing on PA binding proteins. For the purpose of this article, we consider RESPIRATORY BURST OXIDASE HOMOLOGUEs (RBOHs), the most documented PA target proteins. Finally, we present pioneering experiments that show, by different approaches such as monitoring of gene expression, use of pharmacological agents, ectopic over-expression of genes, and the creation of silenced mutants, that lipid signaling plays major roles in crop species. Finally, we present major open questions that require attention since we have only a perception of the peak of the iceberg when it comes to the exciting field of phospholipid signaling in plants.
Keywords: crop species; diacylglycerol kinase; environmental stresses; lipid signaling; phosphatidic acid; phospholipase; protein structure; signaling.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
DIACYLGLYCEROL KINASE 5 participates in flagellin-induced signaling in Arabidopsis.Plant Physiol. 2022 Oct 27;190(3):1978-1996. doi: 10.1093/plphys/kiac354. Plant Physiol. 2022. PMID: 35900211 Free PMC article.
-
Plant phospholipase C family: Regulation and functional role in lipid signaling.Cell Calcium. 2015 Aug;58(2):139-46. doi: 10.1016/j.ceca.2015.04.003. Epub 2015 Apr 17. Cell Calcium. 2015. PMID: 25933832 Review.
-
Plant phosphoinositide-dependent phospholipases C: variations around a canonical theme.Biochimie. 2014 Jan;96:144-57. doi: 10.1016/j.biochi.2013.07.004. Epub 2013 Jul 12. Biochimie. 2014. PMID: 23856562 Review.
-
DIACYLGLYCEROL ACYLTRANSFERASE and DIACYLGLYCEROL KINASE Modulate Triacylglycerol and Phosphatidic Acid Production in the Plant Response to Freezing Stress.Plant Physiol. 2018 Jul;177(3):1303-1318. doi: 10.1104/pp.18.00402. Epub 2018 May 31. Plant Physiol. 2018. PMID: 29853600 Free PMC article.
-
Phospholipase dalpha1 and phosphatidic acid regulate NADPH oxidase activity and production of reactive oxygen species in ABA-mediated stomatal closure in Arabidopsis.Plant Cell. 2009 Aug;21(8):2357-77. doi: 10.1105/tpc.108.062992. Epub 2009 Aug 18. Plant Cell. 2009. PMID: 19690149 Free PMC article.
Cited by
-
Blumeria hordei affects volatile emission of susceptible and resistant barley plants and modifies the defense response of recipient plants.Physiol Plant. 2024 Nov-Dec;176(6):e14646. doi: 10.1111/ppl.14646. Physiol Plant. 2024. PMID: 39648862 Free PMC article.
-
Phosphatidic acid signaling in modulating plant reproduction and architecture.Plant Commun. 2025 Feb 10;6(2):101234. doi: 10.1016/j.xplc.2024.101234. Epub 2024 Dec 24. Plant Commun. 2025. PMID: 39722455 Free PMC article. Review.
References
-
- Furt F., Simon-Plas F., Mongrand S. Lipids of the Plant Plasma Membrane. In: Murphy A.S., Schulz B., Peer W., editors. The Plant Plasma Membrane. Springer; Berlin/Heidelberg, Germany: 2011. pp. 3–30.
LinkOut - more resources
Full Text Sources

