Characterization of Undiscovered miRNA Involved in Tumor Necrosis Factor Alpha-Induced Atrophy in Mouse Skeletal Muscle Cell Line
- PMID: 38892252
- PMCID: PMC11172509
- DOI: 10.3390/ijms25116064
Characterization of Undiscovered miRNA Involved in Tumor Necrosis Factor Alpha-Induced Atrophy in Mouse Skeletal Muscle Cell Line
Abstract
Muscular atrophy is a complex catabolic condition that develops due to several inflammatory-related disorders, resulting in muscle loss. Tumor necrosis factor alpha (TNF-α) is believed to be one of the leading factors that drive inflammatory response and its progression. Until now, the link between inflammation and muscle wasting has been thoroughly investigated, and the non-coding RNA machinery is a potential connection between the candidates. This study aimed to identify specific miRNAs for muscular atrophy induced by TNF-α in the C2C12 murine myotube model. The difference in expression of fourteen known miRNAs and two newly identified miRNAs was recorded by next-generation sequencing between normal muscle cells and treated myotubes. After validation, we confirmed the difference in the expression of one novel murine miRNA (nov-mmu-miRNA-1) under different TNF-α-inducing conditions. Functional bioinformatic analyses of nov-mmu-miRNA-1 revealed the potential association with inflammation and muscle atrophy. Our results suggest that nov-mmu-miRNA-1 may trigger inflammation and muscle wasting by the downregulation of LIN28A/B, an anti-inflammatory factor in the let-7 family. Therefore, TNF-α is involved in muscle atrophy through the modulation of the miRNA cellular machinery. Here, we describe for the first time and propose a mechanism for the newly discovered miRNA, nov-mmu-miRNA-1, which may regulate inflammation and promote muscle atrophy.
Keywords: TNF-α; inflammation; miRNA; muscle atrophy; myotubes.
Conflict of interest statement
The authors have no relevant financial or non-financial interests to disclose.
Figures





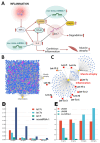
Similar articles
-
TNF-α Induced Myotube Atrophy in C2C12 Cell Line Uncovers Putative Inflammatory-Related lncRNAs Mediating Muscle Wasting.Int J Mol Sci. 2022 Mar 31;23(7):3878. doi: 10.3390/ijms23073878. Int J Mol Sci. 2022. PMID: 35409236 Free PMC article.
-
Decreased miR-497-5p Suppresses IL-6 Induced Atrophy in Muscle Cells.Cells. 2021 Dec 14;10(12):3527. doi: 10.3390/cells10123527. Cells. 2021. PMID: 34944037 Free PMC article.
-
S-allyl cysteine inhibits TNFα-induced skeletal muscle wasting through suppressing proteolysis and expression of inflammatory molecules.Biochim Biophys Acta Gen Subj. 2018 Apr;1862(4):895-906. doi: 10.1016/j.bbagen.2017.12.015. Epub 2017 Dec 28. Biochim Biophys Acta Gen Subj. 2018. PMID: 29288771
-
TWEAK and TRAF6 regulate skeletal muscle atrophy.Curr Opin Clin Nutr Metab Care. 2012 May;15(3):233-9. doi: 10.1097/MCO.0b013e328351c3fc. Curr Opin Clin Nutr Metab Care. 2012. PMID: 22366923 Free PMC article. Review.
-
Muscle wastage in chronic heart failure, between apoptosis, catabolism and altered anabolism: a chimaeric view of inflammation?Curr Opin Clin Nutr Metab Care. 2004 Jul;7(4):435-41. doi: 10.1097/01.mco.0000134374.24181.5b. Curr Opin Clin Nutr Metab Care. 2004. PMID: 15192447 Review.
Cited by
-
TWEAK/Fn14 hypomethylation and higher plasma TWEAK and TNF-α levels are related to sarcopenic obesity in community-dwelling elderly in Xinjiang.Medicine (Baltimore). 2025 Jul 4;104(27):e42937. doi: 10.1097/MD.0000000000042937. Medicine (Baltimore). 2025. PMID: 40629651 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

