MPHGCL-DDI: Meta-Path-Based Heterogeneous Graph Contrastive Learning for Drug-Drug Interaction Prediction
- PMID: 38893359
- PMCID: PMC11173658
- DOI: 10.3390/molecules29112483
MPHGCL-DDI: Meta-Path-Based Heterogeneous Graph Contrastive Learning for Drug-Drug Interaction Prediction
Abstract
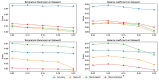
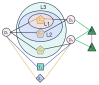
The combinatorial therapy with multiple drugs may lead to unexpected drug-drug interactions (DDIs) and result in adverse reactions to patients. Predicting DDI events can mitigate the potential risks of combinatorial therapy and enhance drug safety. In recent years, deep models based on heterogeneous graph representation learning have attracted widespread interest in DDI event prediction and have yielded satisfactory results, but there is still room for improvement in prediction performance. In this study, we proposed a meta-path-based heterogeneous graph contrastive learning model, MPHGCL-DDI, for DDI event prediction. The model constructs two contrastive views based on meta-paths: an average graph view and an augmented graph view. The former represents that there are connections between drugs, while the latter reveals how the drugs connect with each other. We defined three levels of data augmentation schemes in the augmented graph view and adopted a combination of three losses in the model training phase: multi-relation prediction loss, unsupervised contrastive loss and supervised contrastive loss. Furthermore, the model incorporates indirect drug information, protein-protein interactions (PPIs), to reveal latent relations of drugs. We evaluated MPHGCL-DDI on three different tasks of two datasets. Experimental results demonstrate that MPHGCL-DDI surpasses several state-of-the-art methods in performance.
Keywords: data augmentation; drug-drug interaction; heterogeneous graph contrastive learning; meta-path; protein–protein interaction.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

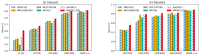
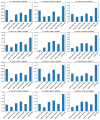
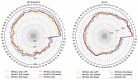
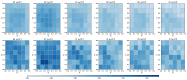

References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

