Advances in Inflammatory Bowel Disease Diagnostics: Machine Learning and Genomic Profiling Reveal Key Biomarkers for Early Detection
- PMID: 38893707
- PMCID: PMC11172026
- DOI: 10.3390/diagnostics14111182
Advances in Inflammatory Bowel Disease Diagnostics: Machine Learning and Genomic Profiling Reveal Key Biomarkers for Early Detection
Abstract
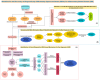
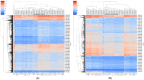
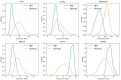
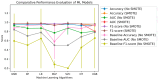
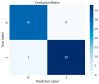
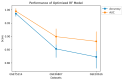
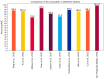
This study, utilizing high-throughput technologies and Machine Learning (ML), has identified gene biomarkers and molecular signatures in Inflammatory Bowel Disease (IBD). We could identify significant upregulated or downregulated genes in IBD patients by comparing gene expression levels in colonic specimens from 172 IBD patients and 22 healthy individuals using the GSE75214 microarray dataset. Our ML techniques and feature selection methods revealed six Differentially Expressed Gene (DEG) biomarkers (VWF, IL1RL1, DENND2B, MMP14, NAAA, and PANK1) with strong diagnostic potential for IBD. The Random Forest (RF) model demonstrated exceptional performance, with accuracy, F1-score, and AUC values exceeding 0.98. Our findings were rigorously validated with independent datasets (GSE36807 and GSE10616), further bolstering their credibility and showing favorable performance metrics (accuracy: 0.841, F1-score: 0.734, AUC: 0.887). Our functional annotation and pathway enrichment analysis provided insights into crucial pathways associated with these dysregulated genes. DENND2B and PANK1 were identified as novel IBD biomarkers, advancing our understanding of the disease. The validation in independent cohorts enhances the reliability of these findings and underscores their potential for early detection and personalized treatment of IBD. Further exploration of these genes is necessary to fully comprehend their roles in IBD pathogenesis and develop improved diagnostic tools and therapies. This study significantly contributes to IBD research with valuable insights, potentially greatly enhancing patient care.
Keywords: differentially expressed genes (DEGs); feature selection (FS); gene ontology; high-throughput technologies; inflammatory bowel disease (IBD); machine learning (ML); pathway enrichment analysis.
Conflict of interest statement
The authors declare that the research was conducted without any commercial or financial relationships construed as a potential conflict of interest.
Figures

References
-
- Alatab S., Sepanlou S.G., Ikuta K., Vahedi H., Bisignano C., Safiri S., Sadeghi A., Nixon M.R., Abdoli A., Abolhassani H., et al. The Global, Regional, and National Burden of Inflammatory Bowel Disease in 195 Countries and Territories, 1990–2017: A Systematic Analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol. Hepatol. 2020;5:17–30. doi: 10.1016/S2468-1253(19)30333-4. - DOI - PMC - PubMed
-
- Wang R., Li Z., Liu S., Zhang D. Global, Regional and National Burden of Inflammatory Bowel Disease in 204 Countries and Territories from 1990 to 2019: A Systematic Analysis Based on the Global Burden of Disease Study 2019. BMJ Open. 2023;13:e065186. doi: 10.1136/bmjopen-2022-065186. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

