Comparative transcriptome and methylome of polar bears, giant and red pandas reveal diet-driven adaptive evolution
- PMID: 38894980
- PMCID: PMC11183199
- DOI: 10.1111/eva.13731
Comparative transcriptome and methylome of polar bears, giant and red pandas reveal diet-driven adaptive evolution
Abstract
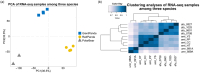
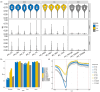
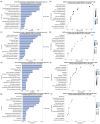
Epigenetic regulation plays an important role in the evolution of species adaptations, yet little information is available on the epigenetic mechanisms underlying the adaptive evolution of bamboo-eating in both giant pandas (Ailuropoda melanoleuca) and red pandas (Ailurus fulgens). To investigate the potential contribution of epigenetic to the adaptive evolution of bamboo-eating in giant and red pandas, we performed hepatic comparative transcriptome and methylome analyses between bamboo-eating pandas and carnivorous polar bears (Ursus maritimus). We found that genes involved in carbohydrate, lipid, amino acid, and protein metabolism showed significant differences in methylation and expression levels between the two panda species and polar bears. Clustering analysis of gene expression revealed that giant pandas did not form a sister group with the more closely related polar bears, suggesting that the expression pattern of genes in livers of giant pandas and red pandas have evolved convergently driven by their similar diets. Compared to polar bears, some key genes involved in carbohydrate metabolism and biological oxidation and cholesterol synthesis showed hypomethylation and higher expression in giant and red pandas, while genes involved in fat digestion and absorption, fatty acid metabolism, lysine degradation, resistance to lipid peroxidation and detoxification showed hypermethylation and low expression. Our study elucidates the special nutrient utilization mechanism of giant pandas and red pandas and provides some insights into the molecular mechanism of their adaptive evolution of bamboo feeding. This has important implications for the breeding and conservation of giant pandas and red pandas.
Keywords: adaptive evolution; giant panda; methylome; polar bear; red panda; transcriptome.
© 2024 The Author(s). Evolutionary Applications published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




Similar articles
-
Gene expressions between obligate bamboo-eating pandas and non-herbivorous mammals reveal converged specialized bamboo diet adaptation.BMC Genomics. 2023 Jan 16;24(1):23. doi: 10.1186/s12864-023-09111-z. BMC Genomics. 2023. PMID: 36647013 Free PMC article.
-
Comparative Transcriptomics and Methylomics Reveal Adaptive Responses of Digestive and Metabolic Genes to Dietary Shift in Giant and Red Pandas.Genes (Basel). 2022 Aug 14;13(8):1446. doi: 10.3390/genes13081446. Genes (Basel). 2022. PMID: 36011357 Free PMC article.
-
Potential Mechanism of Detoxification of Cyanide Compounds by Gut Microbiomes of Bamboo-Eating Pandas.mSphere. 2018 Jun 13;3(3):e00229-18. doi: 10.1128/mSphere.00229-18. Print 2018 Jun 27. mSphere. 2018. PMID: 29898983 Free PMC article.
-
Conservation Genomics and Metagenomics of Giant and Red Pandas in the Wild.Annu Rev Anim Biosci. 2024 Feb 15;12:69-89. doi: 10.1146/annurev-animal-021022-054730. Epub 2023 Oct 20. Annu Rev Anim Biosci. 2024. PMID: 37863091 Review.
-
Giant pandas are not an evolutionary cul-de-sac: evidence from multidisciplinary research.Mol Biol Evol. 2015 Jan;32(1):4-12. doi: 10.1093/molbev/msu278. Epub 2014 Oct 1. Mol Biol Evol. 2015. PMID: 25274274 Review.
Cited by
-
Adaptation in the Alleyways: Candidate Genes Under Potential Selection in Urban Coyotes.Genome Biol Evol. 2025 Jan 6;17(1):evae279. doi: 10.1093/gbe/evae279. Genome Biol Evol. 2025. PMID: 39786569 Free PMC article.
-
Epigenetic Diversity and the Evolutionary Potential of Wild Populations.Evol Appl. 2024 Oct 21;17(10):e70011. doi: 10.1111/eva.70011. eCollection 2024 Oct. Evol Appl. 2024. PMID: 39439434 Free PMC article.
References
-
- Abzhanov, A. , Kuo, W. P. , Hartmann, C. , Grant, B. R. , Grant, P. R. , & Tabin, C. J. (2006). The calmodulin pathway and evolution of elongated beak morphology in Darwin's finches. Nature, 442(7102), 563–567. - PubMed
-
- Bamboo shoots, raw (SR LEGACY, 169210) . (2019). https://fdc.nal.usda.gov/fdc‐app.html#/food‐details/169210/nutrients
LinkOut - more resources
Full Text Sources

