This is a preprint.
Natural variation in protein kinase D modifies alcohol sensitivity in Caenorhabditis elegans
- PMID: 38895441
- PMCID: PMC11185769
- DOI: 10.1101/2024.06.09.598102
Natural variation in protein kinase D modifies alcohol sensitivity in Caenorhabditis elegans
Abstract
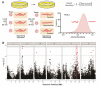
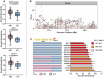
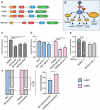
Differences in naïve alcohol sensitivity between individuals are a strong predictor of later life alcohol use disorders (AUD). However, the genetic bases for alcohol sensitivity (beyond ethanol metabolism) and pharmacological approaches to modulate alcohol sensitivity remain poorly understood. We used a high-throughput behavioral screen to measure acute behavioral sensitivity to alcohol, a model of intoxication, in a genetically diverse set of over 150 wild strains of the nematode Caenorhabditis elegans. We performed a genome-wide association study to identify loci that underlie natural variation in alcohol sensitivity. We identified five quantitative trait loci (QTL) and further show that variants in the C. elegans ortholog of protein kinase D, dkf-2, likely underlie the chromosome V QTL. We found that resistance to intoxication was conferred by dkf-2 loss-of-function mutations as well as partly by a PKD inhibitor in a dkf-2-dependent manner. Protein kinase D might represent a conserved, druggable target to modify alcohol sensitivity with application towards AUD.
Keywords: GWAS; alcohol sensitivity; protein kinase D.
Conflict of interest statement
Competing interests: The authors declare no competing interest.
Figures
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
