Viperin-like proteins interfere with RNA viruses in plants
- PMID: 38895613
- PMCID: PMC11185175
- DOI: 10.3389/fpls.2024.1385169
Viperin-like proteins interfere with RNA viruses in plants
Abstract
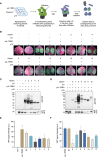
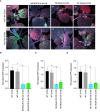
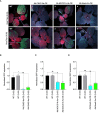
Plant viruses cause substantial losses in crop yield and quality; therefore, devising new, robust strategies to counter viral infections has important implications for agriculture. Virus inhibitory protein endoplasmic reticulum-associated interferon-inducible (Viperin) proteins are conserved antiviral proteins. Here, we identified a set of Viperin and Viperin-like proteins from multiple species and tested whether they could interfere with RNA viruses in planta. Our data from transient and stable overexpression of these proteins in Nicotiana benthamiana reveal varying levels of interference against the RNA viruses tobacco mosaic virus (TMV), turnip mosaic virus (TuMV), and potato virus x (PVX). Harnessing the potential of these proteins represents a novel avenue in plant antiviral approaches, offering a broader and more effective spectrum for application in plant biotechnology and agriculture. Identifying these proteins opens new avenues for engineering a broad range of resistance to protect crop plants against viral pathogens.
Keywords: CNX2; MoaA; RNA virus interference; molybdenum cofactor synthesis 1A (MOCS1A); viperin; viperin-like proteins.
Copyright © 2024 Kamel, Aman and Mahfouz.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Argonaute 5-mediated antiviral defense and viral counter-defense in Nicotiana benthamiana.Virus Res. 2023 Sep;334:199179. doi: 10.1016/j.virusres.2023.199179. Epub 2023 Jul 26. Virus Res. 2023. PMID: 37481165 Free PMC article.
-
Viperin Restricts Zika Virus and Tick-Borne Encephalitis Virus Replication by Targeting NS3 for Proteasomal Degradation.J Virol. 2018 Mar 14;92(7):e02054-17. doi: 10.1128/JVI.02054-17. Print 2018 Apr 1. J Virol. 2018. PMID: 29321318 Free PMC article.
-
MAP30 and luffin-α: Novel ribosome-inactivating proteins induce plant systemic resistance against plant viruses.Pestic Biochem Physiol. 2023 Apr;191:105342. doi: 10.1016/j.pestbp.2023.105342. Epub 2023 Jan 14. Pestic Biochem Physiol. 2023. PMID: 36963924
-
[Progress in studies of viperin, an important cellular antiviral factor].Bing Du Xue Bao. 2015 Jan;31(1):91-6. Bing Du Xue Bao. 2015. PMID: 25997337 Review. Chinese.
-
Molecular biology of resistance to potato virus X in potato.Biochem Soc Symp. 1994;60:207-18. Biochem Soc Symp. 1994. PMID: 7639780 Review.
References
LinkOut - more resources
Full Text Sources

