Revising Clonostachys and allied genera in Bionectriaceae
- PMID: 38895704
- PMCID: PMC11182609
- DOI: 10.3114/sim.2023.105.03
Revising Clonostachys and allied genera in Bionectriaceae
Abstract
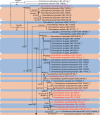
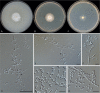
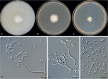
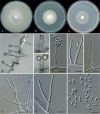
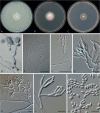
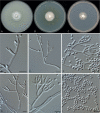
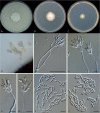
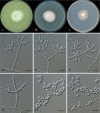
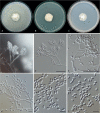
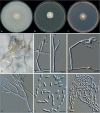
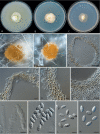
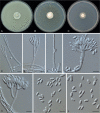
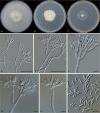
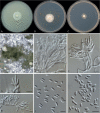
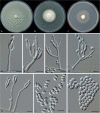
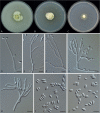
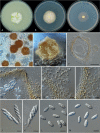
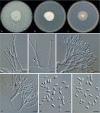
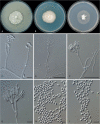
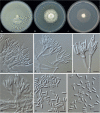
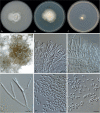
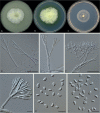
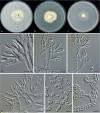
Clonostachys (Bionectriaceae, Hypocreales) species are common soil-borne fungi, endophytes, epiphytes, and saprotrophs. Sexual morphs of Clonostachys spp. were placed in the genus Bionectria, which was further segregated into the six subgenera Astromata, Bionectria, Epiphloea, Myronectria, Uniparietina, and Zebrinella. However, with the end of dual nomenclature, Clonostachys became the single depository for sexual and asexual morph-typified species. Species of Clonostachys are typically characterised by penicillate, sporodochial, and, in many cases, dimorphic conidiophores (primary and secondary conidiophores). Primary conidiophores are mononematous, either verticillium-like or narrowly penicillate. The secondary conidiophores generally form imbricate conidial chains that can collapse to slimy masses, particularly on sporodochia. In the present study, we investigated the species diversity within a collection of 420 strains of Clonostachys from the culture collection of, and personal collections at, the Westerdijk Fungal Biodiversity Institute in Utrecht, the Netherlands. Strains were analysed based on their morphological characters and molecular phylogeny. The latter used DNA sequence data of the nuclear ribosomal internal transcribed spacer regions and intervening 5.8S nrDNA (ITS) and partial 28S large subunit (LSU) nrDNA and partial protein encoding genes including the RNA polymerase II second largest subunit (RPB2), translation elongation factor 1-alpha (TEF1) and β-tubulin (TUB2). Based on these results, the subgenera Astromata, Bionectria, Myronectria and Zebrinella are supported within Clonostachys. Furthermore, the genus Sesquicillium is resurrected to accommodate the former subgenera Epiphloea and Uniparietina. The close relationship of Clonostachys and Sesquicillium is strongly supported as both are inferred phylogenetically as sister-genera. New taxa include 24 new species and 10 new combinations. Recognition of Sesquicillium distinguishes species typically forming a reduced perithecial stroma superficially on plant tissue from species in Clonostachys often forming well-developed, through bark erumpent stromata. The patterns of observed perithecial wall anatomies, perithecial wall and stroma interfaces, and asexual morph diversifications described in a previously compiled monograph are used for interpreting ancestral state reconstructions. It is inferred that the common ancestor of Clonostachys and Sesquicillium may have formed perithecia superficially on leaves, possessed a perithecial wall consisting of a single region, and formed intercalary phialides in penicilli of conidiophores. Character interpretation may also allow hypothesising that diversification of morphs occurred then in the two genera independently and that the frequently stroma-linked Clonostachys morphs evolved together with the occupation of woody host niches and mycoparasitism. Taxonomic novelties: New species: Clonostachys aurantiaca L. Zhao & Crous, Clonostachys australiana L. Zhao & Crous, Clonostachys bambusae L. Zhao & Crous, Clonostachys buxicola L. Zhao & Crous, Clonostachys cylindrica L. Zhao & Crous, Clonostachys ellipsoidea L. Zhao & Crous, Clonostachys flava L. Zhao, Crous & Schroers, Clonostachys fujianensis L. Zhao & Crous, Clonostachys fusca L. Zhao, Crous & Schroers, Clonostachys garysamuelsii L. Zhao & Crous, Clonostachys hongkongensis L. Zhao & Crous, Clonostachys longiphialidica L. Zhao, Crous & Schroers, Clonostachys obovatispora, L. Zhao & Crous, Clonostachys palmae L. Zhao, Crous & Schroers, Clonostachys parasporodochialis L. Zhao & Crous, Clonostachys penicillata L. Zhao, Crous & Schroers, Clonostachys reniformis L. Zhao & Crous, Clonostachys vacuolata L. Zhao, Crous & Schroers, Clonostachys venezuelae L. Zhao, Crous & Schroers, Mycocitrus synnematus L. Zhao & Crous, Nectriopsis didymii L. Zhao & Crous, Sesquicillium intermediophialidicum L. Zhao & Crous, Sesquicillium neerlandicum L. Zhao & Crous, Sesquicillium symmetricum L. Zhao & Crous. New combinations: Mycocitrus coccicola (J.A. Stev.) L. Zhao & Crous, Mycocitrus coxeniae (Y.P. Tan et al.) L. Zhao & Crous, Sesquicillium essexcoheniae (Y.P. Tan et al.) L. Zhao & Crous, Sesquicillium lasiacidis (Samuels) L. Zhao, Crous & Schroers, Sesquicillium phyllophilum (Schroers) L. Zhao, Crous & Schroers, Sesquicillium rossmaniae (Schroers) L. Zhao, Crous & Schroers, Sesquicillium saulense (Lechat & J. Fourn.) L. Zhao & Crous, Sesquicillium sesquicillii (Samuels) L. Zhao, Crous & Schroers, Sesquicillium spinulosisporum (Lechat & J. Fourn.) L. Zhao & Crous, Sesquicillium tornatum (Höhn.) Schroers. New synonyms: Clonostachys aranearum W.H. Chen et al., Clonostachys chuyangsinensis H. Yu & Y. Wang, Clonostachys eriocamporesiana R.H. Perera & K.D. Hyde, Clonostachys granuligera (Starbäck) Forin & Vizzini, Clonostachys indica Prasher & R. Chauhan, Clonostachys spinulosa R.H. Perera et al., Clonostachys squamuligera (Sacc.) Forin & Vizzini, Clonostachys wenpingii (J. Luo & W.Y. Zhuang) Z.Q. Zeng & W.Y. Zhuang. Epitypes (basionyms): Fusidium buxi J.C. Schmidt ex Link, Verticillium candelabrum Bonord. Citation: Zhao L, Groenewald JZ, Hernández-Restrepo M, Schroers H-J, Crous PW (2023). Revising Clonostachys and allied genera in Bionectriaceae. Studies in Mycology 105: 205-266. doi: 10.3114/sim.2023.105.03.
Keywords: Biocontrol; Bionectriaceae; multi-locus; mycoparasitism; new taxa; phylogeny; soil-borne; taxonomy.
© 2023 Westerdijk Fungal Biodiversity Institute.
Conflict of interest statement
The authors declare that there is no conflict of interest.
Figures


















Similar articles
-
Bionectriaceae: a poorly known family of hypocrealean fungi with major commercial potential.Stud Mycol. 2025 Jun;111:115-198. doi: 10.3114/sim.2025.111.04. Epub 2025 Apr 17. Stud Mycol. 2025. PMID: 40371418 Free PMC article.
-
Redisposition of acremonium-like fungi in Hypocreales.Stud Mycol. 2023 Jun;105:23-203. doi: 10.3114/sim.2023.105.02. Epub 2023 Jun 2. Stud Mycol. 2023. PMID: 38895703 Free PMC article.
-
Generic concepts in Nectriaceae.Stud Mycol. 2015 Mar;80:189-245. doi: 10.1016/j.simyco.2014.12.002. Epub 2015 Jan 29. Stud Mycol. 2015. PMID: 26955195 Free PMC article.
-
Fusarium: more than a node or a foot-shaped basal cell.Stud Mycol. 2021 Aug 17;98:100116. doi: 10.1016/j.simyco.2021.100116. eCollection 2021 Apr. Stud Mycol. 2021. PMID: 34466168 Free PMC article.
-
Taxonomy, phylogeny and identification of Chaetomiaceae with emphasis on thermophilic species.Stud Mycol. 2022 Jul;101:121-243. doi: 10.3114/sim.2022.101.03. Epub 2022 Apr 1. Stud Mycol. 2022. PMID: 36059895 Free PMC article.
Cited by
-
Bionectriaceae: a poorly known family of hypocrealean fungi with major commercial potential.Stud Mycol. 2025 Jun;111:115-198. doi: 10.3114/sim.2025.111.04. Epub 2025 Apr 17. Stud Mycol. 2025. PMID: 40371418 Free PMC article.
-
PUPMCR: an R package for image-based identification of color based on Rayner's (1970) terminology and known fungal pigments.Biol Methods Protoc. 2025 Jan 12;10(1):bpaf004. doi: 10.1093/biomethods/bpaf004. eCollection 2025. Biol Methods Protoc. 2025. PMID: 39958703 Free PMC article.
-
Biconidiumsinense gen. et sp. nov. (Hypocreales, Bionectriaceae) and Didymocyrtisshanxiensis sp. nov. (Phaeosphaeriaceae, Didymocyrtis) isolated from urban soil in China.MycoKeys. 2025 Apr 29;116:327-344. doi: 10.3897/mycokeys.116.146683. eCollection 2025. MycoKeys. 2025. PMID: 40337073 Free PMC article.
-
New Tricoderma Species and Six Unrecorded Species of Endolichenic Fungi Isolated from Lichens.Mycobiology. 2025 Jun 23;53(4):435-449. doi: 10.1080/12298093.2025.2519854. eCollection 2025. Mycobiology. 2025. PMID: 40741292 Free PMC article.
-
Eight novel cave fungi in Thailand's Satun Geopark.Fungal Syst Evol. 2023 Nov;12:1-30. doi: 10.3114/fuse.2023.12.01. Epub 2023 Jul 24. Fungal Syst Evol. 2023. PMID: 38455950 Free PMC article.
References
-
- Bainier G. (1905). Mycothèque de l’École de Pharmacie. II. Acrostalagmus roseus Bainier. Bulletin Trimestriel de la Société Mycologique de France 21: 225–228.
-
- Bainier G. (1907). Gliocladium roseum sp. nov. et Cephalosporium acremonium (Corda). Bulletin Trimestriel de la Société Mycologique de France 23: 111–114.
-
- Barnett HL, Lilly VG. (1962). A destructive mycoparasite, Gliocladium roseum. Mycologia 54: 72–77.
-
- Berkeley MJ, Broome CE. (1873). Enumeration of the fungi of Ceylon. Part II. Journal of the Linnean Society. Botany 14: 29–140.
-
- Berkeley MJ, Broome CE. (1876). Notices of British fungi. Annals and Magazine of Natural History. Series 4. 17: 129–145.
LinkOut - more resources
Full Text Sources
Miscellaneous
