Structure and transcription of integrated HPV DNA in vulvar carcinomas
- PMID: 38898085
- PMCID: PMC11187145
- DOI: 10.1038/s41525-024-00418-8
Structure and transcription of integrated HPV DNA in vulvar carcinomas
Abstract
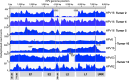
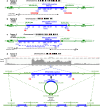
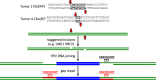
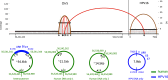
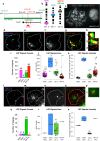
HPV infections are associated with a fraction of vulvar cancers. Through hybridization capture and DNA sequencing, HPV DNA was detected in five of thirteen vulvar cancers. HPV16 DNA was integrated into human DNA in three of the five. The insertions were in introns of human NCKAP1, C5orf67, and LRP1B. Integrations in NCKAP1 and C5orf67 were flanked by short direct repeats in the human DNA, consistent with HPV DNA insertions at sites of abortive, staggered, endonucleolytic incisions. The insertion in C5orf67 was present as a 36 kbp, human-HPV-hetero-catemeric DNA as either an extrachromosomal circle or a tandem repeat within the human genome. The human circularization/repeat junction was defined at single nucleotide resolution. The integrated viral DNA segments all retained an intact upstream regulatory region and the adjacent viral E6 and E7 oncogenes. RNA sequencing revealed that the only HPV genes consistently transcribed from the integrated viral DNAs were E7 and E6*I. The other two HPV DNA+ tumors had coinfections, but no evidence for integration. HPV-positive and HPV-negative vulvar cancers exhibited contrasting human, global gene expression patterns partially overlapping with previously observed differences between HPV-positive and HPV-negative cervical and oropharyngeal cancers. A substantial fraction of the differentially expressed genes involved immune system function. Thus, transcription and HPV DNA integration in vulvar cancers resemble those in other HPV-positive cancers. This study emphasizes the power of hybridization capture coupled with DNA and RNA sequencing to identify a broad spectrum of HPV types, determine human genome integration status of viral DNAs, and elucidate their structures.
© 2024. The Author(s).
Conflict of interest statement
M.H.E. has advised or participated in educational speaking activities but does not receive an honorarium from any companies. In specific cases, his employer has received payment for his time spent on these activities from Asieris, Papivax, Merck, BD, and PDS Biotechnologies. If travel is required for meetings with the industry, the company pays for M.H.E.’s travel expenses. Rutgers has received grant funding for research-related costs of clinical trials that he has been the overall or local PI within the past 12 months from J&J, Pfizer, Inovio, AstraZeneca, Imvax, Iovance, PDS Biotechnologies, and Becton-Dickinson. M.H.E. is a consultant to the World Health Organization. Payment for these efforts goes to Rutgers. M.H.E. has leadership positions in ASCCP, SGO. The other authors declare no potential conflicts of interest.
Figures



References
-
- Surveillance, Epidemiology, and End Results (SEER) Program (www.seer.cancer.gov) SEER*Stat Database: Incidence - SEER Research Data, 8 Registries, Nov 2023 Sub (1975-2021) - Linked To County Attributes - Time Dependent (1990-2022) Income/Rurality, 1969-2022 Counties, National Cancer Institute, DCCPS, Surveillance Research Program, released April 2024, based on the November 2023 submission.
Grants and funding
- R21 CA240580/CA/NCI NIH HHS/United States
- P30 CA072720/CA/NCI NIH HHS/United States
- CA240580/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- P30CA072720/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- P30CA013330P30CA013330/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
LinkOut - more resources
Full Text Sources

