Heterotrimeric Gα-subunit regulates flower and fruit development in CLAVATA signaling pathway in cucumber
- PMID: 38898960
- PMCID: PMC11186068
- DOI: 10.1093/hr/uhae110
Heterotrimeric Gα-subunit regulates flower and fruit development in CLAVATA signaling pathway in cucumber
Abstract
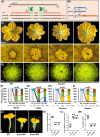
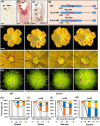
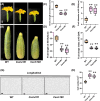
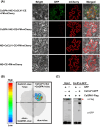
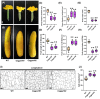
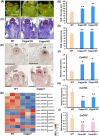
Flowers and fruits are the reproductive organs in plants and play essential roles in natural beauty and the human diet. CLAVATA (CLV) signaling has been well characterized as regulating floral organ development by modulating shoot apical meristem (SAM) size; however, the signaling molecules downstream of the CLV pathway remain largely unknown in crops. Here, we found that functional disruption of CsCLV3 peptide and its receptor CsCLV1 both resulted in flowers with extra organs and stumpy fruits in cucumber. A heterotrimeric G protein α-subunit (CsGPA1) was shown to interact with CsCLV1. Csgpa1 mutant plants derived from gene editing displayed significantly increased floral organ numbers and shorter and wider fruits, a phenotype resembling that of Csclv mutants in cucumber. Moreover, the SAM size was enlarged and the longitudinal cell size of fruit was decreased in Csgpa1 mutants. The expression of the classical stem cell regulator WUSCHEL (WUS) was elevated in the SAM, while the expression of the fruit length stimulator CRABS CLAW (CRC) was reduced in the fruit of Csgpa1 mutants. Therefore, the Gα-subunit CsGPA1 protein interacts with CsCLV1 to inhibit floral organ numbers but promote fruit elongation, via repressing CsWUS expression and activating CsCRC transcription in cucumber. Our findings identified a new player in the CLV signaling pathway during flower and fruit development in dicots, increasing the number of target genes for precise manipulation of fruit shape during crop breeding.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nanjing Agricultural University.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Galli M, Gallavotti A. Expanding the regulatory network for meristem size in plants. Trends Genet. 2016;32:372–83 - PubMed
-
- Ha CM, Jun JH, Fletcher JC. Shoot apical meristem form and function. Curr Top Dev Biol. 2010;91:103–40 - PubMed
-
- Fletcher JC, Brand U, Running MP. et al. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science. 1999;283:1911–4 - PubMed
-
- Clark SE, Williams RW, Meyerowitz EM. The CLAVATA1 gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell. 1997;89:575–85 - PubMed
-
- Ogawa M, Shinohara H, Sakagami Y. et al. Arabidopsis CLV3 peptide directly binds CLV1 ectodomain. Science. 2008;319:294. - PubMed
LinkOut - more resources
Full Text Sources

