Are there double knots in proteins? Prediction and in vitro verification based on TrmD-Tm1570 fusion from C. nitroreducens
- PMID: 38903539
- PMCID: PMC11187310
- DOI: 10.3389/fmolb.2023.1223830
Are there double knots in proteins? Prediction and in vitro verification based on TrmD-Tm1570 fusion from C. nitroreducens
Abstract
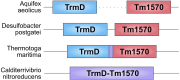
We have been aware of the existence of knotted proteins for over 30 years-but it is hard to predict what is the most complicated knot that can be formed in proteins. Here, we show new and the most complex knotted topologies recorded to date-double trefoil knots (31 #31). We found five domain arrangements (architectures) that result in a doubly knotted structure in almost a thousand proteins. The double knot topology is found in knotted membrane proteins from the CaCA family, that function as ion transporters, in the group of carbonic anhydrases that catalyze the hydration of carbon dioxide, and in the proteins from the SPOUT superfamily that gathers 31 knotted methyltransferases with the active site-forming knot. For each family, we predict the presence of a double knot using AlphaFold and RoseTTaFold structure prediction. In the case of the TrmD-Tm1570 protein, which is a member of SPOUT superfamily, we show that it folds in vitro and is biologically active. Our results show that this protein forms a homodimeric structure and retains the ability to modify tRNA, which is the function of the single-domain TrmD protein. However, how the protein folds and is degraded remains unknown.
Keywords: SPOUT; composite knot; domain; evolution; methyltransferase.
Copyright © 2024 Perlinska, Nguyen, Pilla, Staszor, Lewandowska, Bernat, Purta, Augustyniak, Bujnicki and Sulkowska.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
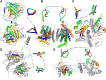



Similar articles
-
Transfer RNA methyltransferases with a SpoU-TrmD (SPOUT) fold and their modified nucleosides in tRNA.Biomolecules. 2017 Feb 28;7(1):23. doi: 10.3390/biom7010023. Biomolecules. 2017. PMID: 28264529 Free PMC article. Review.
-
Structural basis for methyl-donor-dependent and sequence-specific binding to tRNA substrates by knotted methyltransferase TrmD.Proc Natl Acad Sci U S A. 2015 Aug 4;112(31):E4197-205. doi: 10.1073/pnas.1422981112. Epub 2015 Jul 16. Proc Natl Acad Sci U S A. 2015. PMID: 26183229 Free PMC article.
-
Mechanically tightening, untying and retying a protein trefoil knot by single-molecule force spectroscopy.Chem Sci. 2020 Oct 19;11(46):12512-12521. doi: 10.1039/d0sc02796k. Chem Sci. 2020. PMID: 34123232 Free PMC article.
-
AlphaFold predicts the most complex protein knot and composite protein knots.Protein Sci. 2022 Aug;31(8):e4380. doi: 10.1002/pro.4380. Protein Sci. 2022. PMID: 35900026 Free PMC article.
-
[Knotted proteins].Tsitologiia. 2015;57(3):177-83. Tsitologiia. 2015. PMID: 26021166 Review. Russian.
Cited by
-
AlphaKnot 2.0: a web server for the visualization of proteins' knotting and a database of knotted AlphaFold-predicted models.Nucleic Acids Res. 2024 Jul 5;52(W1):W187-W193. doi: 10.1093/nar/gkae443. Nucleic Acids Res. 2024. PMID: 38842945 Free PMC article.
References
LinkOut - more resources
Full Text Sources

