ATM inhibition exploits checkpoint defects and ATM-dependent double strand break repair in TP53-mutant glioblastoma
- PMID: 38906885
- PMCID: PMC11192742
- DOI: 10.1038/s41467-024-49316-8
ATM inhibition exploits checkpoint defects and ATM-dependent double strand break repair in TP53-mutant glioblastoma
Abstract
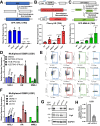
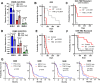
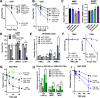
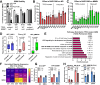
Determining the balance between DNA double strand break repair (DSBR) pathways is essential for understanding treatment response in cancer. We report a method for simultaneously measuring non-homologous end joining (NHEJ), homologous recombination (HR), and microhomology-mediated end joining (MMEJ). Using this method, we show that patient-derived glioblastoma (GBM) samples with acquired temozolomide (TMZ) resistance display elevated HR and MMEJ activity, suggesting that these pathways contribute to treatment resistance. We screen clinically relevant small molecules for DSBR inhibition with the aim of identifying improved GBM combination therapy regimens. We identify the ATM kinase inhibitor, AZD1390, as a potent dual HR/MMEJ inhibitor that suppresses radiation-induced phosphorylation of DSBR proteins, blocks DSB end resection, and enhances the cytotoxic effects of TMZ in treatment-naïve and treatment-resistant GBMs with TP53 mutation. We further show that a combination of G2/M checkpoint deficiency and reliance upon ATM-dependent DSBR renders TP53 mutant GBMs hypersensitive to TMZ/AZD1390 and radiation/AZD1390 combinations. This report identifies ATM-dependent HR and MMEJ as targetable resistance mechanisms in TP53-mutant GBM and establishes an approach for simultaneously measuring multiple DSBR pathways in treatment selection and oncology research.
© 2024. The Author(s).
Conflict of interest statement
J.N.S. reports receiving commercial research grants from AbbVie, ABL Bio, ADC Therapeutics, AstraZeneca, Bayer, Black Diamond, Boehringer Ingelheim, Bristol Myers Squibb, Glaxo Smith Kline, Inhibrx, Karyopharm, ModifiBio, Otomagnetics, Rain Therapeutics, Reglagene, SKBP, Sumitomo Dainippon Pharma Oncology, and Wayshine. Z.D.N. is a co-inventor on a related patent (US 9,938,587 B2) and reports past unrelated sponsored research agreements with Pfizer Inc., Ensoma, Agios, and Intellia Therapeutics.
Figures





References
MeSH terms
Substances
Grants and funding
- T32HL007118/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- P30 ES000002/ES/NIEHS NIH HHS/United States
- U19CA264362/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- U01CA227954/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- P30ES000002/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

