HNF4A and HNF1A exhibit tissue specific target gene regulation in pancreatic beta cells and hepatocytes
- PMID: 38909044
- PMCID: PMC11193738
- DOI: 10.1038/s41467-024-48647-w
HNF4A and HNF1A exhibit tissue specific target gene regulation in pancreatic beta cells and hepatocytes
Abstract
HNF4A and HNF1A encode transcription factors that are important for the development and function of the pancreas and liver. Mutations in both genes have been directly linked to Maturity Onset Diabetes of the Young (MODY) and type 2 diabetes (T2D) risk. To better define the pleiotropic gene regulatory roles of HNF4A and HNF1A, we generated a comprehensive genome-wide map of their binding targets in pancreatic and hepatic cells using ChIP-Seq. HNF4A was found to bind and regulate known (ACY3, HAAO, HNF1A, MAP3K11) and previously unidentified (ABCD3, CDKN2AIP, USH1C, VIL1) loci in a tissue-dependent manner. Functional follow-up highlighted a potential role for HAAO and USH1C as regulators of beta cell function. Unlike the loss-of-function HNF4A/MODY1 variant I271fs, the T2D-associated HNF4A variant (rs1800961) was found to activate AKAP1, GAD2 and HOPX gene expression, potentially due to changes in DNA-binding affinity. We also found HNF1A to bind to and regulate GPR39 expression in beta cells. Overall, our studies provide a rich resource for uncovering downstream molecular targets of HNF4A and HNF1A that may contribute to beta cell or hepatic cell (dys)function, and set up a framework for gene discovery and functional validation.
© 2024. The Author(s).
Conflict of interest statement
N.H.J.N. and A.K.K.T. are co-founders and shareholders of BetaLife Pte Ltd but are not employed by BetaLife Pte Ltd. The remaining authors declare no competing interests.
Figures




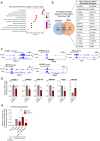



References
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

