Characterization, Structure, and Inhibition of the Human Succinyl-CoA:glutarate-CoA Transferase, a Putative Genetic Modifier of Glutaric Aciduria Type 1
- PMID: 38915184
- PMCID: PMC11259535
- DOI: 10.1021/acschembio.4c00204
Characterization, Structure, and Inhibition of the Human Succinyl-CoA:glutarate-CoA Transferase, a Putative Genetic Modifier of Glutaric Aciduria Type 1
Erratum in
-
Correction to "Characterization, Structure, and Inhibition of the Human Succinyl-CoA:glutarate-CoA Transferase, a Putative Genetic Modifier of Glutaric Aciduria Type 1".ACS Chem Biol. 2025 Nov 21;20(11):2846. doi: 10.1021/acschembio.5c00818. Epub 2025 Oct 23. ACS Chem Biol. 2025. PMID: 41129202 No abstract available.
Abstract
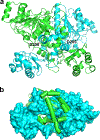
Glutaric Aciduria Type 1 (GA1) is a serious inborn error of metabolism with no pharmacological treatments. A novel strategy to treat this disease is to divert the toxic biochemical intermediates to less toxic or nontoxic metabolites. Here, we report a putative novel target, succinyl-CoA:glutarate-CoA transferase (SUGCT), which we hypothesize suppresses the GA1 metabolic phenotype through decreasing glutaryl-CoA and the derived 3-hydroxyglutaric acid. SUGCT is a type III CoA transferase that uses succinyl-CoA and glutaric acid as substrates. We report the structure of SUGCT, develop enzyme- and cell-based assays, and identify valsartan and losartan carboxylic acid as inhibitors of the enzyme in a high-throughput screen of FDA-approved compounds. The cocrystal structure of SUGCT with losartan carboxylic acid revealed a novel pocket in the active site and further validated the high-throughput screening approach. These results may form the basis for the future development of new pharmacological intervention to treat GA1.
Figures
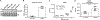



Update of
-
Characterization, structure and inhibition of the human succinyl-CoA:glutarate-CoA transferase, a genetic modifier of glutaric aciduria type 1.bioRxiv [Preprint]. 2024 Feb 7:2024.02.07.578422. doi: 10.1101/2024.02.07.578422. bioRxiv. 2024. Update in: ACS Chem Biol. 2024 Jul 19;19(7):1544-1553. doi: 10.1021/acschembio.4c00204. PMID: 38370847 Free PMC article. Updated. Preprint.
References
-
- Deana R; Rigoni F; Deana AD; Galzigna L, Submitochondrial localization and partial purification of the succinylCoA: 3-hydroxy-3-methylglutarate coenzyme A transferase from rat liver. Biochim Biophys Acta 1981, 662 (1), 119–24. - PubMed
-
- Francesconi MA; Donella-Deana A; Furlanetto V; Cavallini L; Palatini P; Deana R, Further purification and characterization of the succinyl-CoA:3-hydroxy-3-methylglutarate coenzyme A transferase from rat-liver mitochondria. Biochim Biophys Acta 1989, 999 (2), 163–70. - PubMed
-
- Deana R, Substrate specificity of a dicarboxyl-CoA: dicarboxylic acid coenzyme A transferase from rat liver mitochondria. Biochem Int 1992, 26 (4), 767–73. - PubMed
-
- Berthold CL; Toyota CG; Richards NG; Lindqvist Y, Reinvestigation of the catalytic mechanism of formyl-CoA transferase, a class III CoA-transferase. J Biol Chem 2008, 283 (10), 6519–29. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

