Fitness landscape of substrate-adaptive mutations in evolved amino acid-polyamine-organocation transporters
- PMID: 38916596
- PMCID: PMC11198987
- DOI: 10.7554/eLife.93971
Fitness landscape of substrate-adaptive mutations in evolved amino acid-polyamine-organocation transporters
Abstract
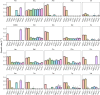
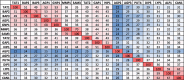
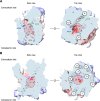
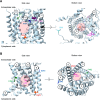
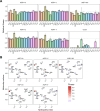
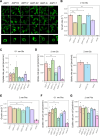
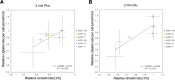
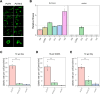
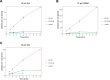
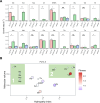
The emergence of new protein functions is crucial for the evolution of organisms. This process has been extensively researched for soluble enzymes, but it is largely unexplored for membrane transporters, even though the ability to acquire new nutrients from a changing environment requires evolvability of transport functions. Here, we demonstrate the importance of environmental pressure in obtaining a new activity or altering a promiscuous activity in members of the amino acid-polyamine-organocation (APC)-type yeast amino acid transporters family. We identify APC members that have broader substrate spectra than previously described. Using in vivo experimental evolution, we evolve two of these transporter genes, AGP1 and PUT4, toward new substrate specificities. Single mutations on these transporters are found to be sufficient for expanding the substrate range of the proteins, while retaining the capacity to transport all original substrates. Nonetheless, each adaptive mutation comes with a distinct effect on the fitness for each of the original substrates, illustrating a trade-off between the ancestral and evolved functions. Collectively, our findings reveal how substrate-adaptive mutations in membrane transporters contribute to fitness and provide insights into how organisms can use transporter evolution to explore new ecological niches.
Keywords: S. cerevisiae; adaptive mutations; amino acid transport; amino acid-polyamine-organocation transport family; biochemistry; chemical biology; evolutionary biology; experimental evolution; yeast.
© 2024, Karapanagioti et al.
Conflict of interest statement
FK, ÚA, DS, BP, SO No competing interests declared
Figures




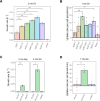
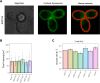
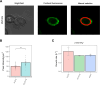
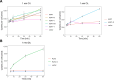



Update of
- doi: 10.1101/2023.10.12.562049
- doi: 10.7554/eLife.93971.1
- doi: 10.7554/eLife.93971.2
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

