Exploring the Subcellular Localization of Monascus Pigments Biosynthases: Preliminary Unraveling of the Compartmentalization Mechanism
- PMID: 38921362
- PMCID: PMC11205011
- DOI: 10.3390/jof10060375
Exploring the Subcellular Localization of Monascus Pigments Biosynthases: Preliminary Unraveling of the Compartmentalization Mechanism
Abstract
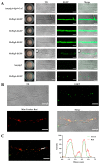
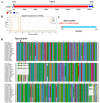
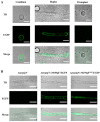
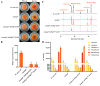
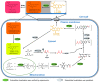
Monascus pigments (MPs), a class of secondary metabolites produced by Monascus spp., can be classified into yellow, orange, and red MPs according to their differences in the wavelength of the maximum absorption. However, the biosynthetic sequence and cellular biosynthesis mechanism of different MPs components are still not yet completely clear in Monascus spp. In this study, the subcellular localization of five MPs synthases was investigated using fluorescent protein fusion expression. The results revealed that the proteins encoded by the MPs biosynthetic gene cluster were compartmentalized in various subcellular locations, including the mitochondrial polyketide synthase MrPigA, cytosolic enzymes consisting of the ketoreductase MrPigC, the oxidoreductase MrPigE, and the monooxygenase MrPigN, and the cell-wall-bound oxidoreductase MrPigF. Moreover, the correct localization of MrPigF to the cell wall was crucial for the synthesis of orange MPs. Lastly, we discussed the compartmentalized biosynthetic pathway of MPs. This study will not only be helpful in clarifying the biosynthetic sequence and biosynthesis mechanism of different MPs but also provides new insights into the cellular biosynthesis of secondary metabolites in filamentous fungi.
Keywords: Monascus pigments; Monascus pigments biosynthetic proteins; Monascus spp.; compartmentalization; subcellular localization.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Dynamic regulation of Monascus azaphilones biosynthesis by the binary MrPigE-MrPigF oxidoreductase system.Appl Microbiol Biotechnol. 2022 Nov;106(22):7519-7530. doi: 10.1007/s00253-022-12219-z. Epub 2022 Oct 12. Appl Microbiol Biotechnol. 2022. PMID: 36221033
-
Comparative transcriptomic analysis reveals the regulatory effects of inorganic nitrogen on the biosynthesis of Monascus pigments and citrinin.RSC Adv. 2020 Feb 3;10(9):5268-5282. doi: 10.1039/c9ra09760k. eCollection 2020 Jan 29. RSC Adv. 2020. PMID: 35498272 Free PMC article.
-
Identification and role analysis of an intermediate produced by a polygenic mutant of Monascus pigments cluster in Monascus ruber M7.Appl Microbiol Biotechnol. 2016 Aug;100(16):7037-49. doi: 10.1007/s00253-016-7397-8. Epub 2016 Mar 5. Appl Microbiol Biotechnol. 2016. PMID: 26946170
-
Production and biological activities of yellow pigments from Monascus fungi.World J Microbiol Biotechnol. 2016 Aug;32(8):136. doi: 10.1007/s11274-016-2082-8. Epub 2016 Jun 29. World J Microbiol Biotechnol. 2016. PMID: 27357404 Review.
-
Monascus pigments.Appl Microbiol Biotechnol. 2012 Dec;96(6):1421-40. doi: 10.1007/s00253-012-4504-3. Epub 2012 Oct 27. Appl Microbiol Biotechnol. 2012. PMID: 23104643 Review.
Cited by
-
Coordinated Synthesis of Pigments Differing in Side Chain Length in Monascus purpureus and Investigation of Pigments and Citrinin Relation.J Agric Food Chem. 2025 Jan 22;73(3):2033-2043. doi: 10.1021/acs.jafc.4c09653. Epub 2025 Jan 10. J Agric Food Chem. 2025. PMID: 39792060 Free PMC article.
-
Purified Monascus Pigments: Biological Activities and Mechanisms of Action.J Nat Prod. 2025 Feb 28;88(2):607-615. doi: 10.1021/acs.jnatprod.4c01008. Epub 2025 Feb 5. J Nat Prod. 2025. PMID: 39906945 Free PMC article. Review.
References
-
- Chen W., He Y., Zhou Y., Shao Y., Feng Y., Li M., Chen F. Edible Filamentous Fungi from the Species Monascus: Early Traditional Fermentations, Modern Molecular Biology, and Future Genomics. Compr. Rev. Food Sci. Food Saf. 2015;14:555–567. doi: 10.1111/1541-4337.12145. - DOI
-
- Li M., Li L., Feng Y., Chen W., He Y., Liu J., Lei M., Liu Q., Shao Y., Chen F. Historic and Charming Monascus spp. Chin. Sci. Bull. 2023;68:479–494. doi: 10.1360/TB-2022-0736. (In Chinese) - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

