Evaluation of RNA Secondary Stem-Loop Structures in the UTRs of Mouse Hepatitis Virus as New Therapeutic Targets
- PMID: 38921815
- PMCID: PMC11206603
- DOI: 10.3390/pathogens13060518
Evaluation of RNA Secondary Stem-Loop Structures in the UTRs of Mouse Hepatitis Virus as New Therapeutic Targets
Abstract
MHV-A59 is a beta-coronavirus that causes demyelinating encephalitis and hepatitis in mice. Recently, the mouse infection model of MHV-A59 has been used as an alternative animal infection model for SARS-CoV and SARS-CoV-2, aiding the development of new antiviral drugs. In this study, the MHV-A59 model was employed to investigate the potential of SARS-CoV-2 UTRs as new targets for antiviral drugs. Optimal targets within the MHV-A59 UTRs were identified using a shRNA and siRNA design tool, focusing on RNA secondary stem-loop (SL) structures in the UTRs. We then examined whether the designed RNAi constructs could inhibit MHV-A59 replication. In the 5'UTR, the stem-loop 1 (SL1) was identified as the most effective target, while in the 3'UTR, the minimal element for the initiation of negative-strand RNA synthesis (MIN) proved to be the most effective. Importantly, siRNAs targeting SL1 and MIN structures significantly reduced total RNA synthesis, negative-strand genomic RNA synthesis, subgenomic (sg) RNA synthesis, viral titer, and the plaque size of MHV-A59 compared to the control. Although not statistically significant, the combination of siSL1 and siMIN had a stronger effect on inhibiting MHV-A59 replication than either siRNA monotherapy. Interestingly, while the SL1 structure is present in both MHV and SARS-CoV-2, the MIN structure is unique to MHV. Thus, the SL1 of SARS-CoV-2 may represent a novel and promising target for RNAi-based antiviral drugs.
Keywords: MHV-A59; UTR; shRNA; siRNA; stem-loop.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
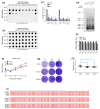






References
-
- Malik Y.A. Properties of coronavirus and SARS-CoV-2. Malays. J. Pathol. 2020;42:3–11. - PubMed
-
- Tan C.W., Chia W.N., Qin X., Liu P., Chen M.I.-C., Tiu C., Hu Z., Chen V.C.-W., Young B.E., Sia W.R. A SARS-CoV-2 surrogate virus neutralization test based on antibody-mediated blockage of ACE2–spike protein–protein interaction. Nat. Biotechnol. 2020;38:1073–1078. doi: 10.1038/s41587-020-0631-z. - DOI - PubMed
-
- Coley S.E., Lavi E., Sawicki S.G., Fu L., Schelle B., Karl N., Siddell S.G., Thiel V. Recombinant mouse hepatitis virus strain A59 from cloned, full-length cDNA replicates to high titers in vitro and is fully pathogenic in vivo. J. Virol. 2005;79:3097–3106. doi: 10.1128/JVI.79.5.3097-3106.2005. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

