Brain Chimeroids reveal individual susceptibility to neurotoxic triggers
- PMID: 38926573
- PMCID: PMC11338177
- DOI: 10.1038/s41586-024-07578-8
Brain Chimeroids reveal individual susceptibility to neurotoxic triggers
Abstract
Interindividual genetic variation affects the susceptibility to and progression of many diseases1,2. However, efforts to study how individual human brains differ in normal development and disease phenotypes are limited by the paucity of faithful cellular human models, and the difficulty of scaling current systems to represent multiple people. Here we present human brain Chimeroids, a highly reproducible, multidonor human brain cortical organoid model generated by the co-development of cells from a panel of individual donors in a single organoid. By reaggregating cells from multiple single-donor organoids at the neural stem cell or neural progenitor cell stage, we generate Chimeroids in which each donor produces all cell lineages of the cerebral cortex, even when using pluripotent stem cell lines with notable growth biases. We used Chimeroids to investigate interindividual variation in the susceptibility to neurotoxic triggers that exhibit high clinical phenotypic variability: ethanol and the antiepileptic drug valproic acid. Individual donors varied in both the penetrance of the effect on target cell types, and the molecular phenotype within each affected cell type. Our results suggest that human genetic background may be an important mediator of neurotoxin susceptibility and introduce Chimeroids as a scalable system for high-throughput investigation of interindividual variation in processes of brain development and disease.
© 2024. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Declaration of interests
P.A. is a SAB member at Foresite Labs, CNSII, and is a co-founder and SAB member of Vesalius Therapeutics. A.R. is a founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas Therapeutics, and until August 31, 2020, was a SAB member of Syros Pharmaceuticals, Neogene Therapeutics, Asimov and Thermo Fisher Scientific. From August 1, 2020, A.R. has been an employee of Genentech and has equity in Roche.
Figures







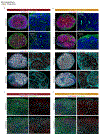

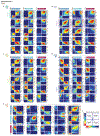








References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

