The Usefulness of qPCR Data for Sample Pre-Assessment and Interpretation of Genetic Typing Results
- PMID: 38927680
- PMCID: PMC11203103
- DOI: 10.3390/genes15060744
The Usefulness of qPCR Data for Sample Pre-Assessment and Interpretation of Genetic Typing Results
Abstract
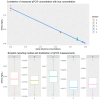
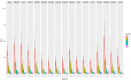
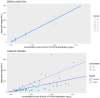
DNA quantification is a crucial step in the STR typing workflow for human identification purposes. Given the reaction's nature, qPCR assays may be subjected to the same stochastic effects of traditional PCR for low-input concentrations. The study aims to evaluate the precision of the PowerQuant® (Promega) kit assay measurements and the degree of variability for DNA templates falling below the optimal threshold of the PowerPlex® ESX-17 Fast STR typing kit (Promega). Five three-fold dilutions of the 2800 M control DNA (Promega) were set up. Each dilution (concentrations: 0.05, 0.0167, 0.0055, 0.00185, and 0.000617 ng/µL) was quantified and amplified in four replicates. Variability for qPCR results, STR profile completeness, and EPGs' peak height were evaluated. The qPCR-estimated concentration of casework samples was correlated with profile completeness and peak intensity, to assess the predictive value of qPCR results for the successful STR typing of scarce samples. qPCR was subjected to stochastic effects, of which the degree was inversely proportional to the initial input template. Quantitation results and the STR profile's characteristics were strongly correlated. Due to the intrinsic nature of real casework samples, a qPCR-derived DNA concentration threshold for correctly identifying probative STR profiles may be difficult to establish. Quantitation data may be useful in interpreting and corroborating STR typing results and for clearly illustrating them to the stakeholders.
Keywords: average peak height; casework samples; forensic genetics; human DNA quantification; probative STR profile; qPCR variation; trace samples.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
-
- Scientific Working Group on DNA Analysis Methods (SWGDAM) Quality Assurance Standards for Forensic DNA Testing Laboratories. Federal Bureau of Investigation; Washington, DC, USA: 2020.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

