Removal and identification of external protein corona members from RBC-derived extracellular vesicles by surface manipulating antimicrobial peptides
- PMID: 38938416
- PMCID: PMC11080927
- DOI: 10.1002/jex2.78
Removal and identification of external protein corona members from RBC-derived extracellular vesicles by surface manipulating antimicrobial peptides
Abstract
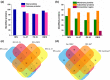
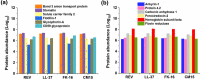
In the last years, extracellular vesicles (EVs), secreted by various cells and body fluids have shown extreme potential in biomedical applications. Increasing number of studies suggest that a protein corona could adhere to the surface of EVs which can have a fundamental effect on their function, targeting and therapeutical efficacy. However, removing and identifying these corona members is currently a challenging task to achieve. In this study we have employed red blood cell-derived extracellular vesicles (REVs) as a model system and three membrane active antimicrobial peptides (AMPs), LL-37, FK-16 and CM15, to test whether they can be used to remove protein corona members from the surface of vesicles. These AMPs were reported to preferentially exert their membrane-related activity via one of the common helical surface-covering models and do not significantly affect the interior of lipid bilayer bodies. The interaction between the peptides and the REVs was followed by biophysical techniques, such as flow-linear dichroism spectroscopy which provided the effective applicable peptide concentration for protein removal. REV samples were then subjected to subsequent size exclusion chromatography and to proteomics analysis. Based on the comparison of control REVs with the peptide treated samples, seventeen proteins were identified as external protein corona members. From the three investigated AMPs, FK-16 can be considered as the best candidate to further optimize EV-related applicability of AMPs. Our results on the REV model system envisage that membrane active peptides may become a useful set of tools in engineering and modifying surfaces of EVs and other lipid-based natural particles.
Keywords: extracellular vesicles; membrane active antimicrobial peptide; protein analysis; protein corona; spectroscopy.
© 2023 The Authors. Journal of Extracellular Biology published by Wiley Periodicals, LLC on behalf of the International Society for Extracellular Vesicles.
Conflict of interest statement
The authors report no conflict of interest.
Figures
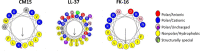



Similar articles
-
Membrane Active Peptides Remove Surface Adsorbed Protein Corona From Extracellular Vesicles of Red Blood Cells.Front Chem. 2020 Aug 11;8:703. doi: 10.3389/fchem.2020.00703. eCollection 2020. Front Chem. 2020. PMID: 32850685 Free PMC article.
-
Categorizing interaction modes of antimicrobial peptides with extracellular vesicles: Disruption, membrane trespassing, and clearance of the protein corona.J Colloid Interface Sci. 2025 Feb;679(Pt A):496-509. doi: 10.1016/j.jcis.2024.09.244. Epub 2024 Oct 1. J Colloid Interface Sci. 2025. PMID: 39378685
-
Biophysical Profiling of Protein Corona on Red Blood Cell-Derived Extracellular Vesicles (REVs): Linear Dichroism and Microfluidic Resistive Pulse Sensing Separate Surface Clearing from Vesicle Disruption.Methods Mol Biol. 2025;2908:213-223. doi: 10.1007/978-1-0716-4434-8_14. Methods Mol Biol. 2025. PMID: 40304912
-
How Membrane-Active Peptides Get into Lipid Membranes.Acc Chem Res. 2016 Jun 21;49(6):1130-8. doi: 10.1021/acs.accounts.6b00074. Epub 2016 May 17. Acc Chem Res. 2016. PMID: 27187572 Review.
-
The Role of Intravesicular Proteins and the Protein Corona of Extracellular Vesicles in the Development of Drug-Induced Polyneuropathy.Curr Issues Mol Biol. 2023 Apr 7;45(4):3302-3314. doi: 10.3390/cimb45040216. Curr Issues Mol Biol. 2023. PMID: 37185740 Free PMC article. Review.
Cited by
-
Factors to consider before choosing EV labeling method for fluorescence-based techniques.Front Bioeng Biotechnol. 2024 Sep 18;12:1479516. doi: 10.3389/fbioe.2024.1479516. eCollection 2024. Front Bioeng Biotechnol. 2024. PMID: 39359260 Free PMC article. Review.
-
A protein corona modulates the function of mineralization-competent matrix vesicles.JBMR Plus. 2024 Dec 24;9(2):ziae168. doi: 10.1093/jbmrpl/ziae168. eCollection 2025 Feb. JBMR Plus. 2024. PMID: 39877729 Free PMC article.
-
In situ captured antibacterial action of membrane-incising peptide lamellae.Nat Commun. 2024 Apr 23;15(1):3424. doi: 10.1038/s41467-024-47708-4. Nat Commun. 2024. PMID: 38654023 Free PMC article.
References
-
- Bebesi, T. , Kitka, D. , Gaál, A. , Szigyártó, I. C. , Deák, R. , Beke‐Somfai, T. , Koprivanacz, K. , Juhász, T. , Bóta, A. , Varga, Z. , & Mihály, J. (2022). Storage conditions determine the characteristics of red blood cell derived extracellular vesicles. Scientific Reports, 12(1), 977. 10.1038/s41598-022-04915-7 - DOI - PMC - PubMed
-
- Bosman, G. J. C. G. M. , Lasonder, E. , Luten, M. , Roerdinkholder‐Stoelwinder, B. , Novotný, V. M. J. , Bos, H. , & De Grip, W. J. (2008). The proteome of red cell membranes and vesicles during storage in blood bank conditions: RBC membrane proteome during storage. Transfusion, 48(5), 827–835. 10.1111/j.1537-2995.2007.01630.x-i2 - DOI - PubMed
LinkOut - more resources
Full Text Sources
