MolPLA: a molecular pretraining framework for learning cores, R-groups and their linker joints
- PMID: 38940143
- PMCID: PMC11211832
- DOI: 10.1093/bioinformatics/btae256
MolPLA: a molecular pretraining framework for learning cores, R-groups and their linker joints
Abstract
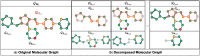
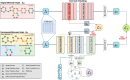
Motivation: Molecular core structures and R-groups are essential concepts in drug development. Integration of these concepts with conventional graph pre-training approaches can promote deeper understanding in molecules. We propose MolPLA, a novel pre-training framework that employs masked graph contrastive learning in understanding the underlying decomposable parts in molecules that implicate their core structure and peripheral R-groups. Furthermore, we formulate an additional framework that grants MolPLA the ability to help chemists find replaceable R-groups in lead optimization scenarios.
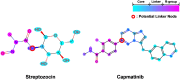
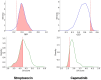
Results: Experimental results on molecular property prediction show that MolPLA exhibits predictability comparable to current state-of-the-art models. Qualitative analysis implicate that MolPLA is capable of distinguishing core and R-group sub-structures, identifying decomposable regions in molecules and contributing to lead optimization scenarios by rationally suggesting R-group replacements given various query core templates.
Availability and implementation: The code implementation for MolPLA and its pre-trained model checkpoint is available at https://github.com/dmis-lab/MolPLA.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures
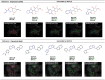
Similar articles
-
MolFCL: predicting molecular properties through chemistry-guided contrastive and prompt learning.Bioinformatics. 2025 Feb 4;41(2):btaf061. doi: 10.1093/bioinformatics/btaf061. Bioinformatics. 2025. PMID: 39921889 Free PMC article.
-
MoleMCL: a multi-level contrastive learning framework for molecular pre-training.Bioinformatics. 2024 Mar 29;40(4):btae164. doi: 10.1093/bioinformatics/btae164. Bioinformatics. 2024. PMID: 38530779 Free PMC article.
-
CMMS-GCL: cross-modality metabolic stability prediction with graph contrastive learning.Bioinformatics. 2023 Aug 1;39(8):btad503. doi: 10.1093/bioinformatics/btad503. Bioinformatics. 2023. PMID: 37572298 Free PMC article.
-
MolMVC: Enhancing molecular representations for drug-related tasks through multi-view contrastive learning.Bioinformatics. 2024 Sep 1;40(Suppl 2):ii190-ii197. doi: 10.1093/bioinformatics/btae386. Bioinformatics. 2024. PMID: 39230706 Free PMC article.
-
Predicting microbe-drug associations with structure-enhanced contrastive learning and self-paced negative sampling strategy.Brief Bioinform. 2023 Mar 19;24(2):bbac634. doi: 10.1093/bib/bbac634. Brief Bioinform. 2023. PMID: 36715986
References
-
- Bemis GW, Murcko MA.. The properties of known drugs. 1. molecular frameworks. J Med Chem 1996;39:2887–93. - PubMed
-
- CTTI. AACT. 2016. https://aact.ctti-clinicaltrials.org/ (17 March 2024, date last accessed).
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

