Stress granule-mediated sequestration of EGR1 mRNAs correlates with lomustine-induced cell death prevention
- PMID: 38940347
- PMCID: PMC11234381
- DOI: 10.1242/jcs.261825
Stress granule-mediated sequestration of EGR1 mRNAs correlates with lomustine-induced cell death prevention
Abstract
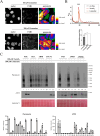
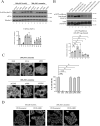
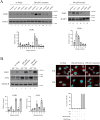
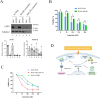
Some chemotherapy drugs modulate the formation of stress granules (SGs), which are RNA-containing cytoplasmic foci contributing to stress response pathways. How SGs mechanistically contribute to pro-survival or pro-apoptotic functions must be better defined. The chemotherapy drug lomustine promotes SG formation by activating the stress-sensing eIF2α kinase HRI (encoded by the EIF2AK1 gene). Here, we applied a DNA microarray-based transcriptome analysis to determine the genes modulated by lomustine-induced stress and suggest roles for SGs in this process. We found that the expression of the pro-apoptotic EGR1 gene was specifically regulated in cells upon lomustine treatment. The appearance of EGR1-encoding mRNA in SGs correlated with a decrease in EGR1 mRNA translation. Specifically, EGR1 mRNA was sequestered to SGs upon lomustine treatment, probably preventing its ribosome translation and consequently limiting the degree of apoptosis. Our data support the model where SGs can selectively sequester specific mRNAs in a stress-specific manner, modulate their availability for translation, and thus determine the fate of a stressed cell.
Keywords: Anticancer drugs; EGR1; Lomustine; Stress granules; Translational control; mRNA translation.
© 2024. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures




References
-
- Adjibade, P., St-Sauveur, V. G., Huberdeau, M. Q., Fournier, M. J., Savard, A., Coudert, L., Khandjian, E. W. and Mazroui, R. (2015). Sorafenib, a multikinase inhibitor, induces formation of stress granules in hepatocarcinoma cells. Oncotarget 6, 43927-43943. 10.18632/oncotarget.5980 - DOI - PMC - PubMed
-
- Adjibade, P., Simoneau, B., Ledoux, N., Gauthier, W. N., Nkurunziza, M., Khandjian, E. W. and Mazroui, R. (2020). Treatment of cancer cells with Lapatinib negatively regulates general translation and induces stress granules formation. PLoS One 15, e0231894. 10.1371/journal.pone.0231894 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

