Taxonomy of introns and the evolution of minor introns
- PMID: 38943346
- PMCID: PMC11347168
- DOI: 10.1093/nar/gkae550
Taxonomy of introns and the evolution of minor introns
Abstract
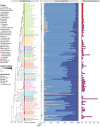
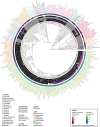
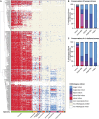
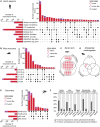
Classification of introns, which is crucial to understanding their evolution and splicing, has historically been binary and has resulted in the naming of major and minor introns that are spliced by their namesake spliceosome. However, a broad range of intron consensus sequences exist, leading us to here reclassify introns as minor, minor-like, hybrid, major-like, major and non-canonical introns in 263 species across six eukaryotic supergroups. Through intron orthology analysis, we discovered that minor-like introns are a transitory node for intron conversion across evolution. Despite close resemblance of their consensus sequences to minor introns, these introns possess an AG dinucleotide at the -1 and -2 position of the 5' splice site, a salient feature of major introns. Through combined analysis of CoLa-seq, CLIP-seq for major and minor spliceosome components, and RNAseq from samples in which the minor spliceosome is inhibited we found that minor-like introns are also an intermediate class from a splicing mechanism perspective. Importantly, this analysis has provided insight into the sequence elements that have evolved to make minor-like introns amenable to recognition by both minor and major spliceosome components. We hope that this revised intron classification provides a new framework to study intron evolution and splicing.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
Minor intron splicing revisited: identification of new minor intron-containing genes and tissue-dependent retention and alternative splicing of minor introns.BMC Genomics. 2019 Aug 30;20(1):686. doi: 10.1186/s12864-019-6046-x. BMC Genomics. 2019. PMID: 31470809 Free PMC article.
-
Origin and evolution of spliceosomal introns.Biol Direct. 2012 Apr 16;7:11. doi: 10.1186/1745-6150-7-11. Biol Direct. 2012. PMID: 22507701 Free PMC article. Review.
-
Roles of minor spliceosome in intron recognition and the convergence with the better understood major spliceosome.Wiley Interdiscip Rev RNA. 2023 Jan;14(1):e1761. doi: 10.1002/wrna.1761. Epub 2022 Sep 2. Wiley Interdiscip Rev RNA. 2023. PMID: 36056453 Review.
-
Evolutionary convergence on highly-conserved 3' intron structures in intron-poor eukaryotes and insights into the ancestral eukaryotic genome.PLoS Genet. 2008 Aug 8;4(8):e1000148. doi: 10.1371/journal.pgen.1000148. PLoS Genet. 2008. PMID: 18688272 Free PMC article.
-
Sequence variations affect the 5' splice site selection of plant introns.Plant Physiol. 2023 Sep 22;193(2):1281-1296. doi: 10.1093/plphys/kiad375. Plant Physiol. 2023. PMID: 37394939
Cited by
-
Pervasive Conservation of Intron Number and Other Genetic Elements Revealed by a Chromosome-level Genome Assembly of the Hyper-polymorphic Nematode Caenorhabditis brenneri.Genome Biol Evol. 2025 Mar 6;17(3):evaf037. doi: 10.1093/gbe/evaf037. Genome Biol Evol. 2025. PMID: 40037811 Free PMC article.
-
Phylogenetic Analysis of 590 Species Reveals Distinct Evolutionary Patterns of Intron-Exon Gene Structures Across Eukaryotic Lineages.Mol Biol Evol. 2024 Dec 6;41(12):msae248. doi: 10.1093/molbev/msae248. Mol Biol Evol. 2024. PMID: 39657604 Free PMC article.
References
-
- Rogozin I.B., Wolf Y.I., Sorokin A.V., Mirkin B.G., Koonin E.V.. Remarkable interkingdom conservation of intron positions and massive, lineage-specific intron loss and gain in eukaryotic evolution. Curr. Biol. 2003; 13:1512–1517. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

