CD4+ T cells display a spectrum of recall dynamics during re-infection with malaria parasites
- PMID: 38944658
- PMCID: PMC11214622
- DOI: 10.1038/s41467-024-49879-6
CD4+ T cells display a spectrum of recall dynamics during re-infection with malaria parasites
Abstract
Children in malaria-endemic regions can experience repeated Plasmodium infections over short periods of time. Effects of re-infection on multiple co-existing CD4+ T cell subsets remain unresolved. Here, we examine antigen-experienced CD4+ T cells during re-infection in mice, using scRNA-seq/TCR-seq and spatial transcriptomics. TCR transgenic TEM cells initiate rapid Th1/Tr1 recall responses prior to proliferating, while GC Tfh counterparts are refractory, with TCM/Tfh-like cells exhibiting modest non-proliferative responses. Th1-recall is a partial facsimile of primary Th1-responses, with no upregulated effector-associated genes being unique to recall. Polyclonal, TCR-diverse, CD4+ T cells exhibit similar recall dynamics, with individual clones giving rise to multiple effectors including highly proliferative Th1/Tr1 cells, as well as GC Tfh and Tfh-like cells lacking proliferative capacity. Thus, we show substantial diversity in recall responses mounted by multiple co-existing CD4+ T cell subsets in the spleen, and present graphical user interfaces for studying gene expression dynamics and clonal relationships during re-infection.
© 2024. The Author(s).
Conflict of interest statement
C.G.W. holds shares in 10x Genomics, whose products were used to generate scRNAseq data in this study. F.C. licensed Slide-seqv2 to Curio Biosciences. No other authors have interests to declare.
Figures





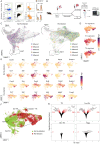


References
-
- World Health Organization, Geneva, Switzerland, Global Malaria Programme. World Malaria Report (World Health Organization, Geneva, Switzerland, 2022).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

