This is a preprint.
Genome-wide association studies of lifetime and frequency cannabis use in 131,895 individuals
- PMID: 38947071
- PMCID: PMC11213095
- DOI: 10.1101/2024.06.14.24308946
Genome-wide association studies of lifetime and frequency cannabis use in 131,895 individuals
Abstract
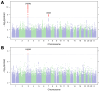
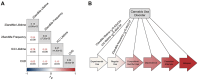
Cannabis is one of the most widely used drugs globally. Decriminalization of cannabis is further increasing cannabis consumption. We performed genome-wide association studies (GWASs) of lifetime (N=131,895) and frequency (N=73,374) of cannabis use. Lifetime cannabis use GWAS identified two loci, one near CADM2 (rs11922956, p=2.40E-11) and another near GRM3 (rs12673181, p=6.90E-09). Frequency of use GWAS identified one locus near CADM2 (rs4856591, p=8.10E-09; r2 =0.76 with rs11922956). Both traits were heritable and genetically correlated with previous GWASs of lifetime use and cannabis use disorder (CUD), as well as other substance use and cognitive traits. Polygenic scores (PGSs) for lifetime and frequency of cannabis use associated cannabis use phenotypes in AllofUs participants. Phenome-wide association study of lifetime cannabis use PGS in a hospital cohort replicated associations with substance use and mood disorders, and uncovered associations with celiac and infectious diseases. This work demonstrates the value of GWASs of CUD transition risk factors.
Keywords: CADM2; GRM3; addiction; cannabis; genetic correlations; genome-wide association study; phenome-wide association study; polygenic score.
Conflict of interest statement
Declaration of Interests PF, the 23andMe Research Team, and S.L.E. are employed by and hold stock or stock options in 23andMe, Inc. The remaining authors have nothing to disclose.
Figures



Similar articles
-
Genome-Wide Association Studies of Impulsive Personality Traits (BIS-11 and UPPS-P) and Drug Experimentation in up to 22,861 Adult Research Participants Identify Loci in the CACNA1I and CADM2 genes.J Neurosci. 2019 Mar 27;39(13):2562-2572. doi: 10.1523/JNEUROSCI.2662-18.2019. Epub 2019 Feb 4. J Neurosci. 2019. PMID: 30718321 Free PMC article.
-
Genome-wide association study of lifetime cannabis use based on a large meta-analytic sample of 32 330 subjects from the International Cannabis Consortium.Transl Psychiatry. 2016 Mar 29;6(3):e769. doi: 10.1038/tp.2016.36. Transl Psychiatry. 2016. PMID: 27023175 Free PMC article.
-
The relationship between cannabis use, schizophrenia, and bipolar disorder: a genetically informed study.Lancet Psychiatry. 2023 Jun;10(6):441-451. doi: 10.1016/S2215-0366(23)00143-8. Lancet Psychiatry. 2023. PMID: 37208114 Free PMC article.
-
The genetic aetiology of cannabis use: from twin models to genome-wide association studies and beyond.Transl Psychiatry. 2022 Nov 21;12(1):489. doi: 10.1038/s41398-022-02215-2. Transl Psychiatry. 2022. PMID: 36411281 Free PMC article. Review.
-
Genetic basis of cannabis use: a systematic review.BMC Med Genomics. 2021 Aug 12;14(1):203. doi: 10.1186/s12920-021-01035-5. BMC Med Genomics. 2021. PMID: 34384432 Free PMC article.
References
-
- United Nations Office on Drug and Crime (2022). Booklet 3 - Drug market trends of Cannabis and Opioids.
-
- Statistics Canada (2013). Mental and substance use disorders in Canada.
Publication types
Grants and funding
- OT2 OD026556/OD/NIH HHS/United States
- R01 DA050721/DA/NIDA NIH HHS/United States
- U2C OD023196/OD/NIH HHS/United States
- OT2 OD025315/OD/NIH HHS/United States
- OT2 OD026551/OD/NIH HHS/United States
- U24 OD023121/OD/NIH HHS/United States
- OT2 OD026552/OD/NIH HHS/United States
- OT2 OD026549/OD/NIH HHS/United States
- OT2 OD025337/OD/NIH HHS/United States
- OT2 OD025277/OD/NIH HHS/United States
- OT2 OD026555/OD/NIH HHS/United States
- K02 AA018755/AA/NIAAA NIH HHS/United States
- OT2 OD023205/OD/NIH HHS/United States
- R01 AA015416/AA/NIAAA NIH HHS/United States
- OT2 OD026554/OD/NIH HHS/United States
- U24 OD023163/OD/NIH HHS/United States
- OT2 OD023206/OD/NIH HHS/United States
- U24 OD023176/OD/NIH HHS/United States
- OT2 OD026548/OD/NIH HHS/United States
- OT2 OD026550/OD/NIH HHS/United States
- P50 AA022537/AA/NIAAA NIH HHS/United States
- DP1 DA054394/DA/NIDA NIH HHS/United States
- OT2 OD026553/OD/NIH HHS/United States
- P50 DA037844/DA/NIDA NIH HHS/United States
- OT2 OD025276/OD/NIH HHS/United States
- U10 AA008401/AA/NIAAA NIH HHS/United States
- OT2 OD026557/OD/NIH HHS/United States
LinkOut - more resources
Full Text Sources
