DUSP1 and SOX2 expression determine squamous cell carcinoma of the salivary gland progression
- PMID: 38951654
- PMCID: PMC11217270
- DOI: 10.1038/s41598-024-65945-x
DUSP1 and SOX2 expression determine squamous cell carcinoma of the salivary gland progression
Abstract
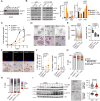
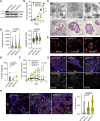
Salivary gland squamous cell carcinomas (SG-SCCs) constitute a rare type of head and neck cancer which is linked to poor prognosis. Due to their low frequency, the molecular mechanisms responsible for their aggressiveness are poorly understood. In this work we studied the role of the phosphatase DUSP1, a negative regulator of MAPK activity, in controlling SG-SCC progression. We generated DUSP1 KO clones in A253 human cells. These clones showed a reduced ability to grow in 2D, self-renew in ECM matrices and to form tumors in immunodeficient mice. This was caused by an overactivation of the stress and apoptosis kinase JNK1/2 in DUSP1-/+ clones. Interestingly, RNAseq analysis revealed that the expression of SOX2, a well-known self-renewal gene was decreased at the mRNA and protein levels in DUSP1-/+ cells. Unexpectedly, CRISPR-KO of SOX2 did not recapitulate DUSP1-/+ phenotype, and SOX2-null cells had an enhanced ability to self-renew and to form tumors in mice. Gene expression analysis demonstrated that SOX2-null cells have a decreased squamous differentiation profile -losing TP63 expression- and an increased migratory phenotype, with an enhanced epithelial to mesenchymal transition signature. In summary, our data indicates that DUSP1 and SOX2 have opposite functions in SG-SCC, being DUSP1 necessary for tumor growth and SOX2 dispensable showing a tumor suppressor function. Our data suggest that the combined expression of SOX2 and DUSP1 could be a useful biomarker to predict progression in patients with SG-SCCs.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
MeSH terms
Substances
Grants and funding
- 23-0272/AICR_/Worldwide Cancer Research/United Kingdom
- PI20/00329/Ministry of Economy and Competitiveness | Instituto de Salud Carlos III (Institute of Health Carlos III)
- CD19/00063/Ministry of Economy and Competitiveness | Instituto de Salud Carlos III (Institute of Health Carlos III)
- PI22/01512/Ministry of Economy and Competitiveness | Instituto de Salud Carlos III (Institute of Health Carlos III)
- PI20/00335/Ministry of Economy and Competitiveness | Instituto de Salud Carlos III (Institute of Health Carlos III)
- PMP22/00083/Ministry of Economy and Competitiveness | Instituto de Salud Carlos III (Institute of Health Carlos III)
- CD23/00167/Ministry of Economy and Competitiveness | Instituto de Salud Carlos III (Institute of Health Carlos III)
- WCRPY224338SAST/Fundación Científica Asociación Española Contra el Cáncer (Scientific Foundation, Spanish Association Against Cancer)
- PEJ-2020-AI/BMD-17846/Comunidad de Madrid
- PI_SEPE_ASP/Comunidad de Madrid
- PEJ-2021-TL_BMD-21631/Comunidad de Madrid
- PEJ-2023-AI/SAL-GL-27700/Comunidad de Madrid
- POSTD20042BEN/Fundación Científica Asociación Española Contra el Cáncer (Scientific Foundation, Spanish Association Against Cancer)
- GLD22/0166/Gilead Sciences (Gilead)
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

