Pan-cancer profiling of tumor-infiltrating natural killer cells through transcriptional reference mapping
- PMID: 38956379
- PMCID: PMC11291284
- DOI: 10.1038/s41590-024-01884-z
Pan-cancer profiling of tumor-infiltrating natural killer cells through transcriptional reference mapping
Abstract
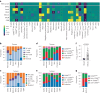
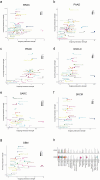
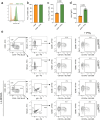
The functional diversity of natural killer (NK) cell repertoires stems from differentiation, homeostatic, receptor-ligand interactions and adaptive-like responses to viral infections. In the present study, we generated a single-cell transcriptional reference map of healthy human blood- and tissue-derived NK cells, with temporal resolution and fate-specific expression of gene-regulatory networks defining NK cell differentiation. Transfer learning facilitated incorporation of tumor-infiltrating NK cell transcriptomes (39 datasets, 7 solid tumors, 427 patients) into the reference map to analyze tumor microenvironment (TME)-induced perturbations. Of the six functionally distinct NK cell states identified, a dysfunctional stressed CD56bright state susceptible to TME-induced immunosuppression and a cytotoxic TME-resistant effector CD56dim state were commonly enriched across tumor types, the ratio of which was predictive of patient outcome in malignant melanoma and osteosarcoma. This resource may inform the design of new NK cell therapies and can be extended through transfer learning to interrogate new datasets from experimental perturbations or disease conditions.
© 2024. The Author(s).
Conflict of interest statement
J.P.G. is an employee at Fate Therapeutics. T.C. is an employee at NEC OncoImmunity AS. K.-J.M. is a consultant at Fate Therapeutics and Vycellix and has research support from Fate Therapeutics, Oncopeptides for studies unrelated to this work. O.D. has received research funding from Gilead Sciences and Incyte, unrelated to this work, and personal fees from Sanofi, unrelated to this work. S.A.T. is a scientific advisory board member of ForeSite Labs, QIAGEN and Element Biosciences, and a co-founder and equity holder of TransitionBio and EnsoCell Therapeutics, and a part-time employee of GlaxoSmithKline. A.H. has received funding from Astra Zeneca/MedImmune, unrelated to this work. A.H. is a consultant at Purple BioTech. The other authors declare no competing interests.
Figures














References
MeSH terms
Substances
Grants and funding
- 2021-073/Ministry of Health and Care Services | Helse Sør-Øst RHF (Southern and Eastern Norway Regional Health Authority)
- 237579/Norges Forskningsråd (Research Council of Norway)
- 275469/Norges Forskningsråd (Research Council of Norway)
- 2021-03069/Vetenskapsrådet (Swedish Research Council)
- 2024-053/Ministry of Health and Care Services | Helse Sør-Øst RHF (Southern and Eastern Norway Regional Health Authority)
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

