Sarcomere, troponin, and myosin X-ray diffraction signals can be resolved in single cardiomyocytes
- PMID: 38956875
- PMCID: PMC11427778
- DOI: 10.1016/j.bpj.2024.06.029
Sarcomere, troponin, and myosin X-ray diffraction signals can be resolved in single cardiomyocytes
Abstract
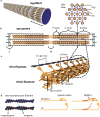
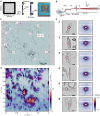
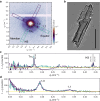
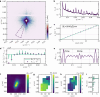
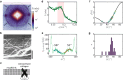
Cardiac function relies on the autonomous molecular contraction mechanisms in the ventricular wall. Contraction is driven by ordered motor proteins acting in parallel to generate a macroscopic force. The averaged structure can be investigated by diffraction from model tissues such as trabecular and papillary cardiac muscle using collimated synchrotron beams, offering high resolution in reciprocal space. In the ventricular wall, however, the muscle tissue is compartmentalized into smaller branched cardiomyocytes, with a higher degree of disorder. We show that X-ray diffraction is now also capable of resolving the structural organization of actomyosin in single isolated cardiomyocytes of the ventricular wall. In addition to the hexagonal arrangement of thick and thin filaments, the diffraction signal of the hydrated and fixated cardiomyocytes was sufficient to reveal the myosin motor repeat (M3), the troponin complex repeat (Tn), and the sarcomere length. The sarcomere length signal comprised up to 13 diffraction orders, which were used to compute the sarcomere density profile based on Fourier synthesis. The Tn and M3 spacings were found in the same range as previously reported for other muscle types. The approach opens up a pathway to record the structural dynamics of living cells during the contraction cycle, toward a more complete understanding of cardiac muscle function.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



Similar articles
-
X-ray diffraction evidence for the extensibility of actin and myosin filaments during muscle contraction.Biophys J. 1994 Dec;67(6):2422-35. doi: 10.1016/S0006-3495(94)80729-5. Biophys J. 1994. PMID: 7779179 Free PMC article.
-
Interference fine structure and sarcomere length dependence of the axial x-ray pattern from active single muscle fibers.Proc Natl Acad Sci U S A. 2000 Jun 20;97(13):7226-31. doi: 10.1073/pnas.97.13.7226. Proc Natl Acad Sci U S A. 2000. PMID: 10860988 Free PMC article.
-
Sarcomere length-dependent effects on Ca2+-troponin regulation in myocardium expressing compliant titin.J Gen Physiol. 2019 Jan 7;151(1):30-41. doi: 10.1085/jgp.201812218. Epub 2018 Dec 6. J Gen Physiol. 2019. PMID: 30523116 Free PMC article.
-
Special Issue: The Actin-Myosin Interaction in Muscle: Background and Overview.Int J Mol Sci. 2019 Nov 14;20(22):5715. doi: 10.3390/ijms20225715. Int J Mol Sci. 2019. PMID: 31739584 Free PMC article. Review.
-
X-ray diffraction studies of the contractile mechanism in single muscle fibres.Philos Trans R Soc Lond B Biol Sci. 2004 Dec 29;359(1452):1883-93. doi: 10.1098/rstb.2004.1557. Philos Trans R Soc Lond B Biol Sci. 2004. PMID: 15647164 Free PMC article. Review.
Cited by
-
Amniotic membrane, a novel bioscaffold in cardiac diseases: from mechanism to applications.Front Bioeng Biotechnol. 2024 Dec 20;12:1521462. doi: 10.3389/fbioe.2024.1521462. eCollection 2024. Front Bioeng Biotechnol. 2024. PMID: 39758951 Free PMC article. Review.
References
-
- Salditt T., Köster S. In: Nanoscale Photonic Imaging. Salditt T., Egner A., Luke D.R., editors. Springer International Publishing; 2020. Scanning Small-Angle X-ray Scattering and Coherent X-ray Imaging of Cells. chapter 15.
-
- Hémonnot C.Y., Köster S. Imaging of biological materials and cells by X-ray scattering and diffraction. ACS Nano. 2017;11:8542–8559. - PubMed
-
- Sakdinawat A., Attwood D. Nanoscale X-ray imaging. Nat. Photonics. 2010;4:840–848.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

