The cortical amygdala consolidates a socially transmitted long-term memory
- PMID: 38961294
- PMCID: PMC11306109
- DOI: 10.1038/s41586-024-07632-5
The cortical amygdala consolidates a socially transmitted long-term memory
Abstract
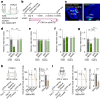
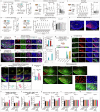
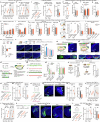
Social communication guides decision-making, which is essential for survival. Social transmission of food preference (STFP) is an ecologically relevant memory paradigm in which an animal learns a desirable food odour from another animal in a social context, creating a long-term memory1,2. How food-preference memory is acquired, consolidated and stored is unclear. Here we show that the posteromedial nucleus of the cortical amygdala (COApm) serves as a computational centre in long-term STFP memory consolidation by integrating social and sensory olfactory inputs. Blocking synaptic signalling by the COApm-based circuit selectively abolished STFP memory consolidation without impairing memory acquisition, storage or recall. COApm-mediated STFP memory consolidation depends on synaptic inputs from the accessory olfactory bulb and on synaptic outputs to the anterior olfactory nucleus. STFP memory consolidation requires protein synthesis, suggesting a gene-expression mechanism. Deep single-cell and spatially resolved transcriptomics revealed robust but distinct gene-expression signatures induced by STFP memory formation in the COApm that are consistent with synapse restructuring. Our data thus define a neural circuit for the consolidation of a socially communicated long-term memory, thereby mechanistically distinguishing protein-synthesis-dependent memory consolidation from memory acquisition, storage or retrieval.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures













Similar articles
-
A Synaptic Circuit Required for Acquisition but Not Recall of Social Transmission of Food Preference.Neuron. 2020 Jul 8;107(1):144-157.e4. doi: 10.1016/j.neuron.2020.04.004. Epub 2020 May 4. Neuron. 2020. PMID: 32369733 Free PMC article.
-
IGF1-Dependent Synaptic Plasticity of Mitral Cells in Olfactory Memory during Social Learning.Neuron. 2017 Jul 5;95(1):106-122.e5. doi: 10.1016/j.neuron.2017.06.015. Neuron. 2017. PMID: 28683263 Free PMC article.
-
Long-term social recognition memory is mediated by oxytocin-dependent synaptic plasticity in the medial amygdala.Biol Psychiatry. 2014 Sep 1;76(5):377-86. doi: 10.1016/j.biopsych.2014.03.022. Epub 2014 Mar 26. Biol Psychiatry. 2014. PMID: 24787950
-
Neural encoding of olfactory recognition memory.J Reprod Dev. 2005 Oct;51(5):547-58. doi: 10.1262/jrd.17031. J Reprod Dev. 2005. PMID: 16284449 Review.
-
The mysteries of remote memory.Philos Trans R Soc Lond B Biol Sci. 2018 Mar 19;373(1742):20170029. doi: 10.1098/rstb.2017.0029. Philos Trans R Soc Lond B Biol Sci. 2018. PMID: 29352028 Free PMC article. Review.
Cited by
-
Cocaine-context memories are transcriptionally encoded in nucleus accumbens Arc ensembles.Nat Commun. 2025 Jul 2;16(1):6084. doi: 10.1038/s41467-025-61004-9. Nat Commun. 2025. PMID: 40603300 Free PMC article.
-
Input-specific properties of excitatory synapses on oxytocin receptor-expressing neurons in the lateral septum.J Neurophysiol. 2024 Dec 1;132(6):1867-1876. doi: 10.1152/jn.00322.2024. Epub 2024 Nov 14. J Neurophysiol. 2024. PMID: 39541198
References
-
- Galef, B. G. Jr. Studies of social learning in Norway rats: a brief review. Dev. Psychobiol.15, 279–295 (1982). - PubMed
-
- Wrenn, C. C. Social transmission of food preference in mice. Curr. Protoc. Neurosci.10.1002/0471142301.ns0805gs28 (2004). - PubMed
-
- Langford, D. J. et al. Social modulation of pain as evidence for empathy in mice. Science312, 1967–1970 (2006). - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

