Altered S-AdenosylMethionine availability impacts dNTP pools in Saccharomyces cerevisiae
- PMID: 38961653
- PMCID: PMC12044341
- DOI: 10.1002/yea.3973
Altered S-AdenosylMethionine availability impacts dNTP pools in Saccharomyces cerevisiae
Abstract
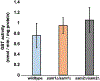
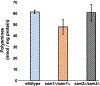
Saccharomyces cerevisiae has long been used as a model organism to study genome instability. The SAM1 and SAM2 genes encode AdoMet synthetases, which generate S-AdenosylMethionine (AdoMet) from Methionine (Met) and ATP. Previous work from our group has shown that deletions of the SAM1 and SAM2 genes cause changes to AdoMet levels and impact genome instability in opposite manners. AdoMet is a key product of methionine metabolism and the major methyl donor for methylation events of proteins, RNAs, small molecules, and lipids. The methyl cycle is interrelated to the folate cycle which is involved in de novo synthesis of purine and pyrimidine deoxyribonucleotides (dATP, dTTP, dCTP, and dGTP). AdoMet also plays a role in polyamine production, essential for cell growth and used in detoxification of reactive oxygen species (ROS) and maintenance of the redox status in cells. This is also impacted by the methyl cycle's role in production of glutathione, another ROS scavenger and cellular protectant. We show here that sam2∆/sam2∆ cells, previously characterized with lower levels of AdoMet and higher genome instability, have a higher level of each dNTP (except dTTP), contributing to a higher overall dNTP pool level when compared to wildtype. Unchecked, these increased levels can lead to multiple types of DNA damage which could account for the genome instability increases in these cells.
Keywords: AdoMet synthetases; Saccharomyces cerevisiae; S‐adenosylmethionine (AdoMet); genomic instability; methyl cycle; yeast genetics.
© 2024 John Wiley & Sons Ltd.
Conflict of interest statement
CONFLICT OF INTEREST STATEMENT
The authors declare no conflict of interest.
Figures

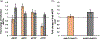
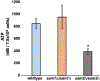

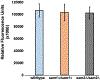
Similar articles
-
Mutations in the S-Adenosylmethionine Synthetase Genes SAM1 and SAM2 Differentially Affect Genome Stability in Saccharomyces cerevisiae.Genetics. 2019 Sep;213(1):97-112. doi: 10.1534/genetics.119.302435. Epub 2019 Jul 18. Genetics. 2019. PMID: 31320408 Free PMC article.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
A nuclear orthologue of the dNTP triphosphohydrolase SAMHD1 controls dNTP homeostasis and genomic stability in Trypanosoma brucei.Front Cell Infect Microbiol. 2023 Aug 22;13:1241305. doi: 10.3389/fcimb.2023.1241305. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37674581 Free PMC article.
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
-
Single-incision sling operations for urinary incontinence in women.Cochrane Database Syst Rev. 2014 Jun 1;(6):CD008709. doi: 10.1002/14651858.CD008709.pub2. Cochrane Database Syst Rev. 2014. Update in: Cochrane Database Syst Rev. 2017 Jul 26;7:CD008709. doi: 10.1002/14651858.CD008709.pub3. PMID: 24880654 Updated.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
