Structures and Functions of the Human GATOR1 Complex
- PMID: 38963491
- PMCID: PMC11997690
- DOI: 10.1007/978-3-031-58843-3_12
Structures and Functions of the Human GATOR1 Complex
Abstract
Eukaryotic cells coordinate available nutrients with their growth through the mechanistic target of rapamycin complex 1 (mTORC1) pathway, in which numerous evolutionarily conserved protein complexes survey and transmit nutrient inputs toward mTORC1. mTORC1 integrates these inputs and activates downstream anabolic or catabolic programs that are in tune with cellular needs, effectively maintaining metabolic homeostasis. The GAP activity toward Rags-1 (GATOR1) protein complex is a critical negative regulator of the mTORC1 pathway and, in the absence of amino acid inputs, is activated to turn off mTORC1 signaling. GATOR1-mediated inhibition of mTORC1 signaling is tightly regulated by an ensemble of protein complexes that antagonize or promote its activity in response to the cellular nutrient environment. Structural, biochemical, and biophysical studies of the GATOR1 complex and its interactors have advanced our understanding of how it regulates cellular metabolism when amino acids are limited. Here, we review the current research with a focus on GATOR1 structure, its enzymatic mechanism, and the growing group of proteins that regulate its activity. Finally, we discuss the implication of GATOR1 dysregulation in physiology and human diseases.
Keywords: Enzyme mechanism; GATOR1; GTPase activating protein (GAP); Protein structure; Rag GTPases; mTORC1.
© 2024. The Author(s), under exclusive license to Springer Nature Switzerland AG.
Figures

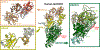


Similar articles
-
Redundant electrostatic interactions between GATOR1 and the Rag GTPase heterodimer drive efficient amino acid sensing in human cells.J Biol Chem. 2023 Jul;299(7):104880. doi: 10.1016/j.jbc.2023.104880. Epub 2023 Jun 1. J Biol Chem. 2023. PMID: 37269949 Free PMC article.
-
Arg-78 of Nprl2 catalyzes GATOR1-stimulated GTP hydrolysis by the Rag GTPases.J Biol Chem. 2019 Feb 22;294(8):2970-2975. doi: 10.1074/jbc.AC119.007382. Epub 2019 Jan 16. J Biol Chem. 2019. PMID: 30651352 Free PMC article.
-
Cryo-EM structures of the human GATOR1-Rag-Ragulator complex reveal a spatial-constraint regulated GAP mechanism.Mol Cell. 2022 May 19;82(10):1836-1849.e5. doi: 10.1016/j.molcel.2022.03.002. Epub 2022 Mar 25. Mol Cell. 2022. PMID: 35338845 Free PMC article.
-
Amino acid-dependent control of mTORC1 signaling: a variety of regulatory modes.J Biomed Sci. 2020 Aug 17;27(1):87. doi: 10.1186/s12929-020-00679-2. J Biomed Sci. 2020. PMID: 32799865 Free PMC article. Review.
-
Regulation of mTORC1 by the Rag GTPases.Biochem Soc Trans. 2023 Apr 26;51(2):655-664. doi: 10.1042/BST20210038. Biochem Soc Trans. 2023. PMID: 36929165 Free PMC article. Review.
Cited by
-
Survival strategies of cancer cells: the role of macropinocytosis in nutrient acquisition, metabolic reprogramming, and therapeutic targeting.Autophagy. 2025 Apr;21(4):693-718. doi: 10.1080/15548627.2025.2452149. Epub 2025 Jan 26. Autophagy. 2025. PMID: 39817564 Free PMC article. Review.
References
-
- Bacq A, Roussel D, Bonduelle T, Zagaglia S, Maletic M, Ribierre T, Adle‐Biassette H, Marchal C, Jennesson M, An I, Picard F, Navarro V, Sisodiya SM, Baulac S, 2022. Cardiac Investigations in Sudden Unexpected Death in DEPDC5 ‐Related Epilepsy. Ann. Neurol. 91, 101–116. 10.1002/ana.26256 - DOI - PMC - PubMed
-
- Bagnall RD, Crompton DE, Petrovski S, Lam L, Cutmore C, Garry SI, Sadleir LG, Dibbens LM, Cairns A, Kivity S, Afawi Z, Regan BM, Duflou J, Berkovic SF, Scheffer IE, Semsarian C, 2016. Exome-based analysis of cardiac arrhythmia, respiratory control, and epilepsy genes in sudden unexpected death in epilepsy. Ann. Neurol. 79, 522–534. 10.1002/ana.24596 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

