Peptide maturation molecules act as molecular gatekeepers to coordinate cell-cell communication in Streptococcus pneumoniae
- PMID: 38963762
- PMCID: PMC11323143
- DOI: 10.1016/j.celrep.2024.114432
Peptide maturation molecules act as molecular gatekeepers to coordinate cell-cell communication in Streptococcus pneumoniae
Abstract
The human pathogen Streptococcus pneumoniae (Spn) encodes several cell-cell communication systems, notably multiple members of the Rgg/SHP and the Tpr/Phr families. Until now, members of these diverse communication systems were thought to work independently. Our study reveals that the ABC transporter PptAB and the transmembrane enzyme Eep act as a molecular link between Rgg/SHP and TprA/PhrA systems. We demonstrate that PptAB/Eep activates the Rgg/SHP systems and represses the TprA/PhrA system. Specifically, they regulate the respective precursor peptides (SHP and PhrA) before these leave the cell. This dual mode of action leads to temporal coordination of these systems, producing an overlap between their respective regulons during host cell infection. Thus, we have identified a single molecular mechanism that targets diverse cell-cell communication systems in Spn. Moreover, these molecular components are encoded by many gram-positive bacteria, suggesting that this mechanism may be broadly conserved.
Keywords: CP: Microbiology; Eep; Gram-positive; PptAB; RRNPPA; Rgg/SHP; Streptococcus pneumoniae; TprA/PhrA; cell-cell communication; colonization; quorum sensing.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures




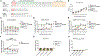
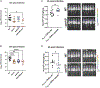
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

