Aberrant evoked calcium signaling and nAChR cluster morphology in a SOD1 D90A hiPSC-derived neuromuscular model
- PMID: 38966427
- PMCID: PMC11222430
- DOI: 10.3389/fcell.2024.1429759
Aberrant evoked calcium signaling and nAChR cluster morphology in a SOD1 D90A hiPSC-derived neuromuscular model
Abstract
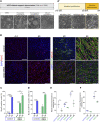
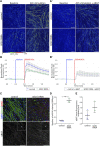
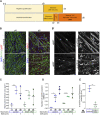
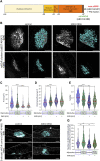
Familial amyotrophic lateral sclerosis (ALS) is a progressive neuromuscular disorder that is due to mutations in one of several target genes, including SOD1. So far, clinical records, rodent studies, and in vitro models have yielded arguments for either a primary motor neuron disease, or a pleiotropic pathogenesis of ALS. While mouse models lack the human origin, in vitro models using human induced pluripotent stem cells (hiPSC) have been recently developed for addressing ALS pathogenesis. In spite of improvements regarding the generation of muscle cells from hiPSC, the degree of maturation of muscle cells resulting from these protocols has remained limited. To fill these shortcomings, we here present a new protocol for an enhanced myotube differentiation from hiPSC with the option of further maturation upon coculture with hiPSC-derived motor neurons. The described model is the first to yield a combination of key myogenic maturation features that are consistent sarcomeric organization in association with complex nAChR clusters in myotubes derived from control hiPSC. In this model, myotubes derived from hiPSC carrying the SOD1 D90A mutation had reduced expression of myogenic markers, lack of sarcomeres, morphologically different nAChR clusters, and an altered nAChR-dependent Ca2+ response compared to control myotubes. Notably, trophic support provided by control hiPSC-derived motor neurons reduced nAChR cluster differences between control and SOD1 D90A myotubes. In summary, a novel hiPSC-derived neuromuscular model yields evidence for both muscle-intrinsic and nerve-dependent aspects of neuromuscular dysfunction in SOD1-based ALS.
Keywords: acetylcholine receptors; amyotrophic lateral sclerosis; hiPSC; motor neurons; myogenesis; neuromuscular junction; skeletal muscle cells; stem cells.
Copyright © 2024 Couturier, Hörner, Nürnberg, Joazeiro, Hafner and Rudolf.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Macrophage-mediated inflammation and glial response in the skeletal muscle of a rat model of familial amyotrophic lateral sclerosis (ALS).Exp Neurol. 2016 Mar;277:275-282. doi: 10.1016/j.expneurol.2016.01.008. Epub 2016 Jan 13. Exp Neurol. 2016. PMID: 26775178 Free PMC article.
-
Coactivation of GSK3β and IGF-1 Attenuates Amyotrophic Lateral Sclerosis Nerve Fiber Cytopathies in SOD1 Mutant Patient-Derived Motor Neurons.Cells. 2021 Oct 16;10(10):2773. doi: 10.3390/cells10102773. Cells. 2021. PMID: 34685754 Free PMC article.
-
Induced pluripotent stem cell-derived motor neurons from amyotrophic lateral sclerosis (ALS) patients carrying different superoxide dismutase 1 mutations recapitulate pathological features of ALS.Chin Med J (Engl). 2021 Oct 6;134(20):2457-2464. doi: 10.1097/CM9.0000000000001693. Chin Med J (Engl). 2021. PMID: 34669638 Free PMC article.
-
Skeletal Muscle Satellite Cells, Mitochondria, and MicroRNAs: Their Involvement in the Pathogenesis of ALS.Front Physiol. 2016 Sep 13;7:403. doi: 10.3389/fphys.2016.00403. eCollection 2016. Front Physiol. 2016. PMID: 27679581 Free PMC article. Review.
-
Mutant SOD1 mediated pathogenesis of Amyotrophic Lateral Sclerosis.Gene. 2016 Feb 15;577(2):109-18. doi: 10.1016/j.gene.2015.11.049. Epub 2015 Dec 2. Gene. 2016. PMID: 26657039 Review.
References
-
- Adler M., Manley H. A., Purcell A. L., Deshpande S. S., Hamilton T. A., Kan R. K., et al. (2004). Reduced acetylcholine receptor density, morphological remodeling, and butyrylcholinesterase activity can sustain muscle function in acetylcholinesterase knockout mice. Muscle Nerve 30, 317–327. 10.1002/mus.20099 - DOI - PubMed
-
- Ahlers J., Althviz Moré D., Amsalem O., Anderson A., Bokota G., Boone P., et al. (2023). napari: a multi-dimensional image viewer for Python. Zenodo. 10.5281/zenodo.3555620 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

