Computational Insights into Amide Bond Formation Catalyzed by the Condensation Domain of Nonribosomal Peptide Synthetases
- PMID: 38973878
- PMCID: PMC11223147
- DOI: 10.1021/acsomega.4c02531
Computational Insights into Amide Bond Formation Catalyzed by the Condensation Domain of Nonribosomal Peptide Synthetases
Abstract
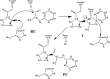
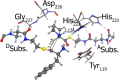
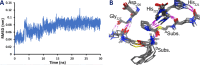
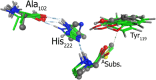
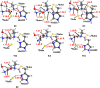
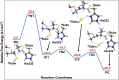
Nonribosomal peptide synthetases (NRPSs) are important enzymes that synthesize an array of nongenetically encoded peptides. The latter have diverse physicochemical properties and roles. NRPSs are modular enzymes in which, for example, the condensation (C-) domain catalyzes the formation of amide bonds. The NRPS tyrocidine synthetase from Brevibacillus brevis is responsible for synthesizing the cyclic-peptide antibiotic tyrocidine. The first step is formation of an amide bond between a proline and phenylalanine which is catalyzed by a C-domain. In this study, a multiscale computational approach (molecular dynamics and QM/MM) has been used to investigate substrate binding and catalytic mechanism of the C-domain of tyrocidine synthetase. Overall, the mechanism is found to proceed through three exergonic steps in which an active site Histidine, His222, acts as a base and acid. First, His222 acts as a base to facilitate nucleophilic attack of the prolyl nitrogen at the phenylalanyl's carbonyl carbon. This is also the rate-limiting step with a free energy barrier of 38.8 kJ mol-1. The second step is collapse of the resulting tetrahedral intermediate with cleavage of the S-C bond between the phenylalanyl and its Ppant arm, along with formation of the above amide bond. Meanwhile, the now protonated His222 imidazole has rotated toward the newly formed thiolate of the Ppant arm. In the final step, His222 acts as an acid, protonating the thiolate and regenerating a neutral His222. The overall mechanism is found to be exergonic with the final product complex being 46.3 kJ mol-1 lower in energy than the initial reactant complex.
© 2024 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
Similar articles
-
Structural and mutational analysis of the nonribosomal peptide synthetase heterocyclization domain provides insight into catalysis.Proc Natl Acad Sci U S A. 2017 Jan 3;114(1):95-100. doi: 10.1073/pnas.1614191114. Epub 2016 Dec 19. Proc Natl Acad Sci U S A. 2017. PMID: 27994138 Free PMC article.
-
Backbone resonance assignments of the C-terminal thioesterase domain of tyrocidine synthetase C.Biomol NMR Assign. 2025 Jun;19(1):17-21. doi: 10.1007/s12104-024-10210-5. Epub 2024 Dec 11. Biomol NMR Assign. 2025. PMID: 39661265
-
Peptide mixed phosphonates for covalent complex formation with thioesterases in nonribosomal peptide synthetases.J Pept Sci. 2024 Jan;30(1):e3532. doi: 10.1002/psc.3532. Epub 2023 Jul 9. J Pept Sci. 2024. PMID: 37423887
-
Structural and functional aspects of the nonribosomal peptide synthetase condensation domain superfamily: discovery, dissection and diversity.Biochim Biophys Acta Proteins Proteom. 2017 Nov;1865(11 Pt B):1587-1604. doi: 10.1016/j.bbapap.2017.05.010. Epub 2017 May 16. Biochim Biophys Acta Proteins Proteom. 2017. PMID: 28526268 Review.
-
Biosynthesis of depsipeptides, or Depsi: The peptides with varied generations.Protein Sci. 2020 Dec;29(12):2316-2347. doi: 10.1002/pro.3979. Epub 2020 Nov 2. Protein Sci. 2020. PMID: 33073901 Free PMC article. Review.
Cited by
-
Structure, Function and Engineering of the Nonribosomal Peptide Synthetase Condensation Domain.Int J Mol Sci. 2024 Nov 1;25(21):11774. doi: 10.3390/ijms252111774. Int J Mol Sci. 2024. PMID: 39519324 Free PMC article. Review.
-
Structures and mechanism of condensation in non-ribosomal peptide synthesis.Nature. 2025 Feb;638(8049):270-278. doi: 10.1038/s41586-024-08417-6. Epub 2024 Dec 11. Nature. 2025. PMID: 39662504
References
-
- Matilla M. A.; Krell T.. Plant Growth Promotion and Biocontrol Mediated by Plant-Associated Bacteria. In Plant Microbiome: Stress Response; Egamberdieva D., Ahmad P., Eds.; Springer Singapore: Singapore, 2018; pp 45–80.
-
- Giordano D.; Coppola D.; Russo R.; Denaro R.; Giuliano L.; Lauro F. M.; di Prisco G.; Verde C.. Chapter Four—Marine Microbial Secondary Metabolites: Pathways, Evolution and Physiological Roles. In Advances in Microbial Physiology; Poole R. K., Ed.; Academic Press, 2015; Vol. 66, pp 357–428. - PubMed
LinkOut - more resources
Full Text Sources
