Genomic and phenotypic inconsistencies in Pseudomonas aeruginosa resistome among intensive care patients
- PMID: 38975326
- PMCID: PMC11224958
- DOI: 10.3389/fcimb.2024.1335096
Genomic and phenotypic inconsistencies in Pseudomonas aeruginosa resistome among intensive care patients
Abstract
Objective: Pseudomonas aeruginosa, a difficult-to-manage nosocomial pathogen, poses a serious threat to clinical outcomes in intensive care (ICU) patients due to its high antimicrobial resistance (AMR). To promote effective management, it is essential to investigate the genomic and phenotypic differences in AMR expression of the isolates.
Methods: A prospective observational study was conducted from July 2022 to April 2023 at Liepaja Regional Hospital in Latvia. The study included all adult patients who were admitted to the ICU and had a documented infection with P. aeruginosa, as confirmed by standard laboratory microbiological testing and short-read sequencing. Since ResFinder is the only sequencing-based database offering antibacterial susceptibility testing (AST) data for each antibiotic, we conducted a comparison of the resistance profile with the results of phenotypic testing, evaluating if ResFinder met the US Food and Drug Administration (FDA) requirements for approval as a new AMR diagnostic test. Next, to improve precision, AST data from ResFinder was compared with two other databases - AMRFinderPlus and RGI. Additionally, data was gathered from environmental samples to inform the implementation of appropriate infection control measures in real time.
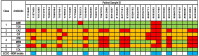
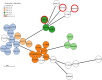
Results: Our cohort consisted of 33 samples from 29 ICU patients and 34 environmental samples. The presence of P. aeruginosa infection was found to be associated with unfavourable clinical outcomes. A third of the patient samples were identified as multi-drug resistant isolates. Apart from resistance against colistin, significant discrepancies were observed when phenotypic data were compared to genotypic data. For example, the aminoglycoside resistance prediction of ResFinder yielded a major errors value of 3.03% for amikacin, which was marginally above the FDA threshold. Among the three positive environmental samples, one sample exhibited multiple AMR genes similar to the patient samples in its cluster.
Conclusion: Our findings underscore the importance of utilizing a combination of diagnostic methods for the identification of resistance mechanisms, clusters, and environmental reservoirs in ICUs.
Keywords: Pseudomonas aeruginosa; antimicrobial resistance; antimicrobial susceptibility testing; environmental reservoirs; intensive care unit; next-generation sequencing.
Copyright © 2024 Dolgusevs, Jain, Savicka, Vangravs, Bodrenko, Bergmanis, Zemite, Selderina, Reinis and Rozentale.
Conflict of interest statement
All authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Alcock B. P., Huynh W., Chalil R., Smith K. W., Raphenya A. R., Wlodarski M. A., et al. . (2023). CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 51, D690–D699. doi: 10.1093/nar/gkac920 - DOI - PMC - PubMed
-
- Ananda T., Vandana K. E., Mukhopadhyay C. (2024). Comparative evaluation of Vitek®2 and broth microdilution method for colistin susceptibility testing of Gram-negative isolates from intensive care unit in a tertiary care hospital. Indian J. Med. Microbiol. 48, 100559. doi: 10.1016/j.ijmmb.2024.100559 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

