A diagnostic model for Parkinson's disease based on circadian rhythm-related genes
- PMID: 38978048
- PMCID: PMC11229228
- DOI: 10.1186/s12967-024-05424-z
A diagnostic model for Parkinson's disease based on circadian rhythm-related genes
Abstract
Background: Circadian rhythm (CR) disturbance is intricately associated with Parkinson's disease (PD). However, the involvement of CR-related mechanisms in the pathogenesis and progression of PD remains elusive.
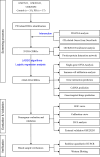
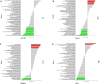
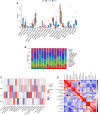
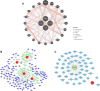
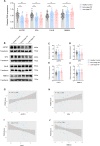
Methods: A total of 141 PD patients and 113 healthy participants completed CR-related clinical examinations in this study. To further investigate the CR-related mechanisms in PD, we obtained datasets (GSE7621, GSE20141, GSE20292) from the Gene Expression Omnibus database to identify differentially expressed genes between PD patients and healthy controls and further selected CR-related genes (CRRGs). Subsequently, the least absolute shrinkage and selection operator (LASSO) followed by logistic algorithms were employed to identify the hub genes and construct a diagnostic model. The predictive performance was evaluated by area under the curve (AUC), calibration curve, and decision curve analyses in the training set and external validation sets. Finally, RT‒qPCR and Western blotting were conducted to verify the expression of these hub genes in blood samples. In addition, Pearson correlation analysis was utilized to validate the association between expression of hub genes and circadian rhythm function.
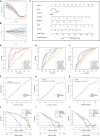
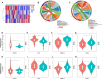
Results: Our clinical observational study revealed that even early-stage PD patients exhibited a higher likelihood of experiencing sleep disturbances, nocturnal hypertension, reverse-dipper blood pressure, and reduced heart rate variability compared to healthy controls. Furthermore, 4 CR-related hub genes (AGTR1, CALR, BRM14, and XPA) were identified and subsequently incorporated as candidate biomarkers to construct a diagnostic model. The model showed satisfactory diagnostic performance in the training set (AUC = 0.941), an external validation set GSE20295 (AUC = 0.842), and our clinical centre set (AUC = 0.805). Additionally, the up-regulation of CALR, BRM14 and the down-regulation of AGTR1, XPA were associated with circadian rhythm disruption.
Conclusion: CR disturbance seems to occur in the early stage of PD. The diagnostic model based on CR-related genes demonstrated robust diagnostic efficacy, offering novel insights for future clinical diagnosis of PD and providing a foundation for further exploration into the role of CR-related mechanisms in the progression of PD.
Keywords: Bioinformatics; Circadian rhythm; Nomogram; Parkinson’s disease.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

