Clinicopathological analysis of NEK1 variants in amyotrophic lateral sclerosis
- PMID: 38986433
- PMCID: PMC11669413
- DOI: 10.1111/bpa.13287
Clinicopathological analysis of NEK1 variants in amyotrophic lateral sclerosis
Abstract
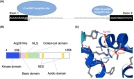
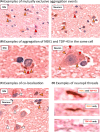
Many genes have been linked to amyotrophic lateral sclerosis (ALS), including never in mitosis A (NIMA)-related kinase 1 (NEK1), a serine/threonine kinase that plays a key role in several cellular functions, such as DNA damage response and cell cycle regulation. Whole-exome sequencing studies have shown that NEK1 mutations are associated with an increased risk for ALS, where a significant enrichment of NEK1 loss-of-function (LOF) variants were found in individuals with ALS compared to controls. In particular, the p.Arg261His missense variant was associated with significantly increased disease susceptibility. This case series aims to understand the neuropathological phenotypes resulting from NEK1 mutations in ALS. We examined a cohort of three Scottish patients with a mutation in the NEK1 gene and evaluated the distribution and cellular expression of NEK1, as well as the abundance of phosphorylated TDP-43 (pTDP-43) aggregates, in the motor cortex compared to age- and sex-matched control tissue. We show pathological, cytoplasmic TDP-43 aggregates in all three NEK1-ALS cases. NEK1 protein staining revealed no immunoreactivity in two of the NEK1-ALS cases, indicating a LOF and corresponding to a reduction in NEK1 mRNA as detected by in situ hybridisation. However, the p.Arg261His missense mutation resulted in an increase in NEK1 mRNA molecules and abundant NEK1-positive cytoplasmic aggregates, with the same morphologic appearance, and within the same cells as co-occurring TDP-43 aggregates. Here we show the first neuropathological assessment of a series of ALS cases carrying mutations in the NEK1 gene. Specifically, we show that TDP-43 pathology is present in these cases and that potential NEK1 LOF can either be mediated through loss of NEK1 translation or through aggregation of NEK1 protein as in the case with p.Arg261His mutation, a potential novel pathological feature of NEK1-ALS.
Keywords: ALS; NEK1; genetics; neuropathology; post‐mortem.
© 2024 The Author(s). Brain Pathology published by John Wiley & Sons Ltd on behalf of International Society of Neuropathology.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

