Emerging methylation-based approaches in microbiome engineering
- PMID: 38987811
- PMCID: PMC11238421
- DOI: 10.1186/s13068-024-02529-x
Emerging methylation-based approaches in microbiome engineering
Abstract
Bacterial epigenetics, particularly through DNA methylation, exerts significant influence over various biological processes such as DNA replication, uptake, and gene regulation in bacteria. In this review, we explore recent advances in characterizing bacterial epigenomes, accompanied by emerging strategies that harness bacterial epigenetics to elucidate and engineer diverse bacterial species with precision and effectiveness. Furthermore, we delve into the potential of epigenetic modifications to steer microbial functions and influence community dynamics, offering promising opportunities for understanding and modulating microbiomes. Additionally, we investigate the extensive diversity of DNA methyltransferases and emphasize their potential utility in the context of the human microbiome. In summary, this review highlights the potential of DNA methylation as a powerful toolkit for engineering microbiomes.
Keywords: Bacterial epigenetics; DNA methyltransferases; Methylome; Microbiome engineering; Restriction-modification (R-M) systems.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


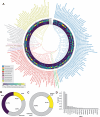
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
