PEPhub: a database, web interface, and API for editing, sharing, and validating biological sample metadata
- PMID: 38991851
- PMCID: PMC11238423
- DOI: 10.1093/gigascience/giae033
PEPhub: a database, web interface, and API for editing, sharing, and validating biological sample metadata
Abstract
Background: As biological data increase, we need additional infrastructure to share them and promote interoperability. While major effort has been put into sharing data, relatively less emphasis is placed on sharing metadata. Yet, sharing metadata is also important and in some ways has a wider scope than sharing data themselves.
Results: Here, we present PEPhub, an approach to improve sharing and interoperability of biological metadata. PEPhub provides an API, natural-language search, and user-friendly web-based sharing and editing of sample metadata tables. We used PEPhub to process more than 100,000 published biological research projects and index them with fast semantic natural-language search. PEPhub thus provides a fast and user-friendly way to finding existing biological research data or to share new data.
Availability: https://pephub.databio.org.
Keywords: metadata API; metadata machine learning; metadata sharing; metadata validation.
© The Author(s) 2024. Published by Oxford University Press GigaScience.
Conflict of interest statement
N.C.S. is a consultant for InVitro Cell Research, LLC. All other authors declare no competing interests.
Figures


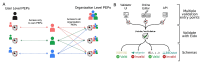
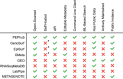
Update of
-
PEPhub: a database, web interface, and API for editing, sharing, and validating biological sample metadata.bioRxiv [Preprint]. 2024 May 11:2023.08.15.551388. doi: 10.1101/2023.08.15.551388. bioRxiv. 2024. Update in: Gigascience. 2024 Jan 2;13:giae033. doi: 10.1093/gigascience/giae033. PMID: 37645717 Free PMC article. Updated. Preprint.
Similar articles
-
PEPhub: a database, web interface, and API for editing, sharing, and validating biological sample metadata.bioRxiv [Preprint]. 2024 May 11:2023.08.15.551388. doi: 10.1101/2023.08.15.551388. bioRxiv. 2024. Update in: Gigascience. 2024 Jan 2;13:giae033. doi: 10.1093/gigascience/giae033. PMID: 37645717 Free PMC article. Updated. Preprint.
-
MetaBasis: a web-based database containing metadata on software tools and databases in the field of bioinformatics.Appl Bioinformatics. 2006;5(3):187-92. doi: 10.2165/00822942-200605030-00007. Appl Bioinformatics. 2006. PMID: 16922600
-
A digital repository with an extensible data model for biobanking and genomic analysis management.BMC Genomics. 2014;15 Suppl 3(Suppl 3):S3. doi: 10.1186/1471-2164-15-S3-S3. Epub 2014 May 6. BMC Genomics. 2014. PMID: 25077808 Free PMC article.
-
Web tools for predictive toxicology model building.Expert Opin Drug Metab Toxicol. 2012 Jul;8(7):791-801. doi: 10.1517/17425255.2012.685158. Epub 2012 May 12. Expert Opin Drug Metab Toxicol. 2012. PMID: 22577953 Review.
-
Evolution of web services in bioinformatics.Brief Bioinform. 2005 Jun;6(2):178-88. doi: 10.1093/bib/6.2.178. Brief Bioinform. 2005. PMID: 15975226 Review.
Cited by
-
Using semantic search to find publicly available gene-expression datasets.bioRxiv [Preprint]. 2025 Mar 15:2025.03.13.643153. doi: 10.1101/2025.03.13.643153. bioRxiv. 2025. PMID: 40161731 Free PMC article. Preprint.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

