Turn-on protein switches for controlling actin binding in cells
- PMID: 38992021
- PMCID: PMC11239668
- DOI: 10.1038/s41467-024-49934-2
Turn-on protein switches for controlling actin binding in cells
Abstract
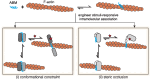
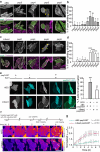
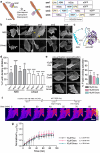
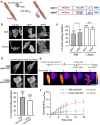
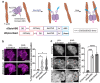
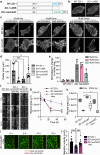
Within a shared cytoplasm, filamentous actin (F-actin) plays numerous and critical roles across the cell body. Cells rely on actin-binding proteins (ABPs) to organize F-actin and to integrate its polymeric characteristics into diverse cellular processes. Yet, the multitude of ABPs that engage with and shape F-actin make studying a single ABP's influence on cellular activities a significant challenge. Moreover, without a means of manipulating actin-binding subcellularly, harnessing the F-actin cytoskeleton for synthetic biology purposes remains elusive. Here, we describe a suite of designed proteins, Controllable Actin-binding Switch Tools (CASTs), whose actin-binding behavior can be controlled with external stimuli. CASTs were developed that respond to different external inputs, providing options for turn-on kinetics and enabling orthogonality and multiplexing. Being genetically encoded, we show that CASTs can be inserted into native protein sequences to control F-actin association locally and engineered into structures to control cell and tissue shape and behavior.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Update of
-
Turn-On Protein Switches for Controlling Actin Binding in Cells.bioRxiv [Preprint]. 2023 Oct 26:2023.10.26.561921. doi: 10.1101/2023.10.26.561921. bioRxiv. 2023. Update in: Nat Commun. 2024 Jul 11;15(1):5840. doi: 10.1038/s41467-024-49934-2. PMID: 37961502 Free PMC article. Updated. Preprint.
Similar articles
-
Turn-On Protein Switches for Controlling Actin Binding in Cells.bioRxiv [Preprint]. 2023 Oct 26:2023.10.26.561921. doi: 10.1101/2023.10.26.561921. bioRxiv. 2023. Update in: Nat Commun. 2024 Jul 11;15(1):5840. doi: 10.1038/s41467-024-49934-2. PMID: 37961502 Free PMC article. Updated. Preprint.
-
F-Actin Cytoskeleton Network Self-Organization Through Competition and Cooperation.Annu Rev Cell Dev Biol. 2020 Oct 6;36:35-60. doi: 10.1146/annurev-cellbio-032320-094706. Annu Rev Cell Dev Biol. 2020. PMID: 33021819 Free PMC article. Review.
-
Regulation of actin cytoskeleton dynamics in cells.Mol Cells. 2010 Apr;29(4):311-25. doi: 10.1007/s10059-010-0053-8. Mol Cells. 2010. PMID: 20446344 Free PMC article. Review.
-
Cytosolic concentrations of actin binding proteins and the implications for in vivo F-actin turnover.J Cell Biol. 2023 Dec 4;222(12):e202306036. doi: 10.1083/jcb.202306036. Epub 2023 Oct 6. J Cell Biol. 2023. PMID: 37801069 Free PMC article.
-
Exploring the Role of the Plant Actin Cytoskeleton: From Signaling to Cellular Functions.Int J Mol Sci. 2023 Oct 23;24(20):15480. doi: 10.3390/ijms242015480. Int J Mol Sci. 2023. PMID: 37895158 Free PMC article. Review.
Cited by
-
Structure of the F-tractin-F-actin complex.J Cell Biol. 2025 Apr 7;224(4):e202409192. doi: 10.1083/jcb.202409192. Epub 2025 Feb 10. J Cell Biol. 2025. PMID: 39928047 Free PMC article.
-
Binding memory of liquid molecules.Nat Commun. 2025 Jul 16;16(1):6555. doi: 10.1038/s41467-025-61630-3. Nat Commun. 2025. PMID: 40670366 Free PMC article.
References
-
- Mehidi A, et al. Forces generated by lamellipodial actin filament elongation regulate the WAVE complex during cell migration. Nat. Chem. Biol. 2021;23:1148–1162. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

