GolpHCat (TMEM87A), a unique voltage-dependent cation channel in Golgi apparatus, contributes to Golgi-pH maintenance and hippocampus-dependent memory
- PMID: 38992057
- PMCID: PMC11239671
- DOI: 10.1038/s41467-024-49297-8
GolpHCat (TMEM87A), a unique voltage-dependent cation channel in Golgi apparatus, contributes to Golgi-pH maintenance and hippocampus-dependent memory
Abstract
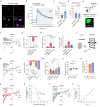
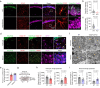
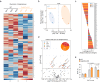
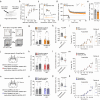
Impaired ion channels regulating Golgi pH lead to structural alterations in the Golgi apparatus, such as fragmentation, which is found, along with cognitive impairment, in Alzheimer's disease. However, the causal relationship between altered Golgi structure and cognitive impairment remains elusive due to the lack of understanding of ion channels in the Golgi apparatus of brain cells. Here, we identify that a transmembrane protein TMEM87A, renamed Golgi-pH-regulating cation channel (GolpHCat), expressed in astrocytes and neurons that contributes to hippocampus-dependent memory. We find that GolpHCat displays unique voltage-dependent currents, which is potently inhibited by gluconate. Additionally, we gain structural insights into the ion conduction through GolpHCat at the molecular level by determining three high-resolution cryogenic-electron microscopy structures of human GolpHCat. GolpHCat-knockout mice show fragmented Golgi morphology and altered protein glycosylation and functions in the hippocampus, leading to impaired spatial memory. These findings suggest a molecular target for Golgi-related diseases and cognitive impairment.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
miR-142 downregulation alleviates the impairment of spatial learning and memory, reduces the level of apoptosis, and upregulates the expression of pCaMKII and BAI3 in the hippocampus of APP/PS1 transgenic mice.Behav Brain Res. 2021 Sep 24;414:113485. doi: 10.1016/j.bbr.2021.113485. Epub 2021 Jul 21. Behav Brain Res. 2021. PMID: 34302879
-
Transmembrane proteins with unknown function (TMEMs) as ion channels: electrophysiological properties, structure, and pathophysiological roles.Exp Mol Med. 2024 Apr;56(4):850-860. doi: 10.1038/s12276-024-01206-1. Epub 2024 Apr 1. Exp Mol Med. 2024. PMID: 38556553 Free PMC article. Review.
-
Neuronal and astroglial monocarboxylate transporters play key but distinct roles in hippocampus-dependent learning and memory formation.Prog Neurobiol. 2020 Nov;194:101888. doi: 10.1016/j.pneurobio.2020.101888. Epub 2020 Jul 18. Prog Neurobiol. 2020. PMID: 32693190
-
Structure of the GOLD-domain seven-transmembrane helix protein family member TMEM87A.Elife. 2022 Nov 14;11:e81704. doi: 10.7554/eLife.81704. Elife. 2022. PMID: 36373655 Free PMC article.
-
[pH homeostasis of the Golgi apparatus by GPHR, a novel anion channel].Nihon Yakurigaku Zasshi. 2011 May;137(5):212-6. doi: 10.1254/fpj.137.212. Nihon Yakurigaku Zasshi. 2011. PMID: 21558671 Review. Japanese. No abstract available.
Cited by
-
Golgi pH homeostasis stabilizes the lysosomal membrane through N-glycosylation of membrane proteins.Life Sci Alliance. 2024 Jul 30;7(10):e202402677. doi: 10.26508/lsa.202402677. Print 2024 Oct. Life Sci Alliance. 2024. PMID: 39079741 Free PMC article.
-
Acute GARP Depletion Disrupts Vesicle Transport, Leading to Severe Defects in Sorting, Secretion and O-Glycosylation.Traffic. 2025 Jan-Mar;26(1-3):e70003. doi: 10.1111/tra.70003. Traffic. 2025. PMID: 40100055 Free PMC article.
-
The mechanosensitive channel ELKIN1 regulates cellular adaptations to simulated microgravity.NPJ Microgravity. 2025 Mar 16;11(1):10. doi: 10.1038/s41526-025-00466-z. NPJ Microgravity. 2025. PMID: 40090965 Free PMC article.
-
Acute GARP depletion disrupts vesicle transport, leading to severe defects in sorting, secretion, and O-glycosylation.bioRxiv [Preprint]. 2024 Oct 14:2024.10.07.617053. doi: 10.1101/2024.10.07.617053. bioRxiv. 2024. Update in: Traffic. 2025 Jan-Mar;26(1-3):e70003. doi: 10.1111/tra.70003. PMID: 39416116 Free PMC article. Updated. Preprint.
References
-
- Maeda, Y. & Kinoshita, T. J. M. The acidic environment of the Golgi is critical for glycosylation and transport. Methods Enzymol. 480, 495–510 (2010). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

