Identification and functional analysis of long non-coding RNA (lncRNA) and metabolites response to mowing in hulless barley (Hordeum vulgare L. var. nudum hook. f.)
- PMID: 38997634
- PMCID: PMC11241897
- DOI: 10.1186/s12870-024-05334-8
Identification and functional analysis of long non-coding RNA (lncRNA) and metabolites response to mowing in hulless barley (Hordeum vulgare L. var. nudum hook. f.)
Abstract
Background: Hulless barley (Hordeum vulgare L. var. nudum Hook. f.) is a significant cereal crop and a substantial source of forage for livestock. Long non-coding RNAs (lncRNAs) and metabolites play crucial roles in the nutrient accumulation and regeneration of hulless barley plants following mowing. The study aimed to identify differentially expressed lncRNAs and metabolites in hulless barley plants by analyzing transcriptomic and metabolomic datasets at 2 h, 24 h, and 72 h following mowing.
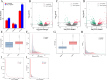
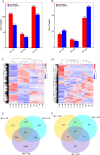
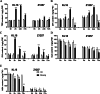
Results: The study revealed that 190, 90, and 438 lncRNA genes were differentially expressed at the 2 h, 24 h, and 72 h time points compared to the non-mowing control. We identified 14 lncRNA genes-11 downregulated and 3 upregulated-showing consistently significant differential expression across all time points after mowing. These differentially expressed lncRNAs target genes involved in critical processes such as cytokinin signaling, cell wall degradation, storage protein accumulation, and biomass increase. In addition, we identified ten differentially expressed metabolites targeting diverse metabolic pathways, including plant hormones, alkaloids, and flavonoids, before and after mowing at various time points. Endogenous hormone analysis revealed that cytokinin most likely played a crucial role in the regeneration of hulless barley after mowing.
Conclusions: This study created a comprehensive dataset of lncRNAs, metabolites, and hormones in hulless barley after mowing, revealing valuable insights into the functional characteristics of lncRNAs, metabolites, and hormones in regulating plant regeneration. The results indicated that cytokinin plays a significant role in facilitating the regeneration process of hulless barley after mowing. This comprehensive dataset is an invaluable resource for better understanding the complex mechanisms that underlie plant regeneration, with significant implications for crop improvement.
Keywords: Cytokinin; Hulless barley; Metabolome; Mowing; Plant hormones; lncRNA.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Transcriptome Screening of Long Noncoding RNAs and Their Target Protein-Coding Genes Unmasks a Dynamic Portrait of Seed Coat Coloration Associated with Anthocyanins in Tibetan Hulless Barley.Int J Mol Sci. 2023 Jun 24;24(13):10587. doi: 10.3390/ijms241310587. Int J Mol Sci. 2023. PMID: 37445765 Free PMC article.
-
Accumulation and regulation of anthocyanins in white and purple Tibetan Hulless Barley (Hordeum vulgare L. var. nudum Hook. f.) revealed by combined de novo transcriptomics and metabolomics.BMC Plant Biol. 2022 Aug 4;22(1):391. doi: 10.1186/s12870-022-03699-2. BMC Plant Biol. 2022. PMID: 35922757 Free PMC article.
-
Identification of HvLRX, a new dehydration and light responsive gene in Tibetan hulless barley (Hordeum vulgare var. nudum).Genes Genomics. 2021 Dec;43(12):1445-1461. doi: 10.1007/s13258-021-01147-3. Epub 2021 Sep 3. Genes Genomics. 2021. PMID: 34480266
-
Dehydration induced transcriptomic responses in two Tibetan hulless barley (Hordeum vulgare var. nudum) accessions distinguished by drought tolerance.BMC Genomics. 2017 Oct 11;18(1):775. doi: 10.1186/s12864-017-4152-1. BMC Genomics. 2017. PMID: 29020945 Free PMC article.
-
Underground communication: Long non-coding RNA signaling in the plant rhizosphere.Plant Commun. 2024 Jul 8;5(7):100927. doi: 10.1016/j.xplc.2024.100927. Epub 2024 Apr 27. Plant Commun. 2024. PMID: 38679911 Free PMC article. Review.
Cited by
-
Comparative transcriptome analysis reveals key long noncoding RNAs for cadmium tolerance in Tibetan hull-less barley.Front Plant Sci. 2025 May 22;16:1572490. doi: 10.3389/fpls.2025.1572490. eCollection 2025. Front Plant Sci. 2025. PMID: 40475905 Free PMC article.
-
A long non-coding RNA lncRNA18313 regulates resistance against cadmium stress in wheat.Front Plant Sci. 2025 Jun 2;16:1583758. doi: 10.3389/fpls.2025.1583758. eCollection 2025. Front Plant Sci. 2025. PMID: 40530269 Free PMC article.
References
-
- Guo T, Horvath C, Chen L, Chen J, Zheng B. Understanding the nutrient composition and nutritional functions of highland barley (Qingke): a review. Trends Food Sci Technol. 2020;103:109–117. doi: 10.1016/j.tifs.2020.07.011. - DOI
-
- WU K-L, Yao X-H, Yao Y-H, Bai Y-X, Chi D-Z. Reflections and practice on breeding barley varieties under the background of diversified uses. Tibet J Agri Sci. 2018;40(S1):1–2.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

