SlMDH3 Interacts with Autophagy Receptor Protein SlATI1 and Positively Regulates Tomato Heat Tolerance
- PMID: 39000108
- PMCID: PMC11241746
- DOI: 10.3390/ijms25137000
SlMDH3 Interacts with Autophagy Receptor Protein SlATI1 and Positively Regulates Tomato Heat Tolerance
Abstract
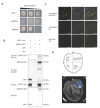
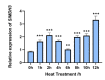
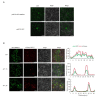
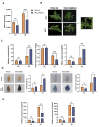
Autophagy, a highly conserved protein degradation system, plays an important role in protecting cells from adverse environmental conditions. ATG8-INTERACTING PROTEIN1 (ATI1) acts as an autophagy receptor, but its functional mechanisms in plants' heat stress tolerance remain unclear. In this study, using LC-MS/MS, we identified malate dehydrogenase (SlMDH3) as a SlATI1-interacting protein. Further studies showed that heat stress induced the expression of SlMDH3 and SlMDH3 co-localized with SlATI1 under both 22 °C and 42 °C heat treatment conditions. Moreover, silencing of SlMDH3 increased the sensitivity of tomato to heat stress, as evidenced by exacerbated degradation of chlorophyll; accumulation of MDA, H2O2, and dead cells; increased relative conductivity; and inhibition of stress-related gene expression. Conversely, overexpression of SlMDH3 improved tomato's heat tolerance, leading to opposite effects on physiological indicators and gene expression compared to SlMDH3 silencing. Taken together, our study suggests that SlMDH3 interacts with SlATI1 and positively regulates tomato heat tolerance.
Keywords: ATI1; autophagy; heat stress; malate dehydrogenase; tomato.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Overexpression of SlATG8f gene enhanced autophagy and pollen protection in tomato under heat stress.Sci Rep. 2024 Nov 6;14(1):26892. doi: 10.1038/s41598-024-77491-7. Sci Rep. 2024. PMID: 39505980 Free PMC article.
-
Overexpression of Solanum habrochaites microRNA319d (sha-miR319d) confers chilling and heat stress tolerance in tomato (S. lycopersicum).BMC Plant Biol. 2019 May 23;19(1):214. doi: 10.1186/s12870-019-1823-x. BMC Plant Biol. 2019. PMID: 31122194 Free PMC article.
-
WHIRLY1 Regulates HSP21.5A Expression to Promote Thermotolerance in Tomato.Plant Cell Physiol. 2020 Jan 1;61(1):169-177. doi: 10.1093/pcp/pcz189. Plant Cell Physiol. 2020. PMID: 31596474
-
Interactions between 2-Cys peroxiredoxins and ascorbate in autophagosome formation during the heat stress response in Solanum lycopersicum.J Exp Bot. 2016 Mar;67(6):1919-33. doi: 10.1093/jxb/erw013. Epub 2016 Jan 31. J Exp Bot. 2016. PMID: 26834179 Free PMC article.
-
Involvement of HSP70 in BAG9-mediated thermotolerance in Solanum lycopersicum.Plant Physiol Biochem. 2024 Feb;207:108353. doi: 10.1016/j.plaphy.2024.108353. Epub 2024 Jan 9. Plant Physiol Biochem. 2024. PMID: 38219426
References
-
- Hemantaranjan A., Bhanu A.N., Singh M.N., Yadav D.K., Patel P.K., Singh R., Katiyar D. Heat Stress Responses and Thermotolerance. Adv. Plants Agric. Res. 2014;1:1–10. doi: 10.15406/apar.2014.01.00012. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

