This is a preprint.
Deciphering the cis-regulatory landscape of natural yeast Transcript Leaders
- PMID: 39005336
- PMCID: PMC11245039
- DOI: 10.1101/2024.07.03.601937
Deciphering the cis-regulatory landscape of natural yeast Transcript Leaders
Update in
-
Deciphering the landscape of cis-acting sequences in natural yeast transcript leaders.Nucleic Acids Res. 2025 Feb 27;53(5):gkaf165. doi: 10.1093/nar/gkaf165. Nucleic Acids Res. 2025. PMID: 40071932 Free PMC article.
Abstract
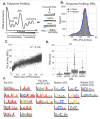
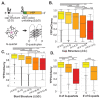
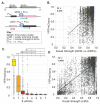
Protein synthesis is a vital process that is highly regulated at the initiation step of translation. Eukaryotic 5' transcript leaders (TLs) contain a variety of cis-regulatory features that influence translation and mRNA stability. However, the relative influences of these features in natural TLs are poorly characterized. To address this, we used massively parallel reporter assays (MPRAs) to quantify RNA levels, ribosome loading, and protein levels from 11,027 natural yeast TLs in vivo and systematically compared the relative impacts of their sequence features on gene expression. We found that yeast TLs influence gene expression over two orders of magnitude. While a leaky scanning model using Kozak contexts and uAUGs explained half of the variance in expression across transcript leaders, the addition of other features explained ~70% of gene expression variation. Our analyses detected key cis-acting sequence features, quantified their effects in vivo, and compared their roles to motifs reported from an in vitro study of ribosome recruitment. In addition, our work quantitated the effects of alternative transcription start site usage on gene expression in yeast. Thus, our study provides new quantitative insights into the roles of TL cis-acting sequences in regulating gene expression.
Figures



References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
