This is a preprint.
Bulk and Single-cell Transcriptomic Brain Data Identify Overlapping Processes and Cell-types with Human AUD and Mammalian Models of Alcohol Use
- PMID: 39005451
- PMCID: PMC11245012
- DOI: 10.1101/2024.07.02.601528
Bulk and Single-cell Transcriptomic Brain Data Identify Overlapping Processes and Cell-types with Human AUD and Mammalian Models of Alcohol Use
Abstract
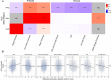
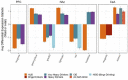
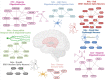
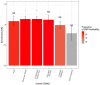
This study explores the neurobiological underpinnings of alcohol use disorder (AUD) by integrating bulk and single-cell transcriptomic data from humans, primates, and mice across three brain regions associated with addiction (i.e., prefrontal cortex (PFC), nucleus accumbens (NAc), and central amygdala (CeA)). We compared AUD RNA expression and cell-type abundance from 92 human brain to data from 53 primates and 90 mice engaged in diverse alcohol use paradigms. The findings revealed significant and reproducible correlations between human AUD and mammalian models of alcohol use that vary by tissue, species, and behavioral paradigm. The strongest correlations occurred between primate and mouse models of binge drinking (i.e., high drinking in the dark). Certain primate models demonstrated that the brain RNA correlations with human alcohol use disorder (AUD) were approximately 40% as strong as the correlations observed within human samples themselves. By integrating single-cell transcriptomic data, this study observed decreased oligodendrocyte proportions in the PFC and NAc of human AUD with similar trends in animal models. Gene co-expression network analyses revealed conserved systems associated with human AUD and animal models of heavy/binge alcohol consumption. Gene co-expression networks were enriched for pathways related to inflammation, myelination, and synaptic plasticity and the genes within them accounted for ~20% of the heritability in human alcohol consumption. Identified hub genes were associated with relevant traits (e.g., impulsivity, motivation) in humans and mice. This study sheds light on conserved biological entities underlying AUD and chronic alcohol use, providing insights into the cellular, genetic, and neuromolecular basis across species.
Conflict of interest statement
Conflicts: The authors have no conflicts to declare.
Figures

References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
