Amphibian Segmentation Clock Models Suggest How Large Genome and Cell Sizes Slow Developmental Rate
- PMID: 39006893
- PMCID: PMC11245677
- DOI: 10.1093/iob/obae021
Amphibian Segmentation Clock Models Suggest How Large Genome and Cell Sizes Slow Developmental Rate
Abstract
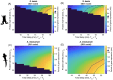
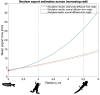
Evolutionary increases in genome size, cell volume, and nuclear volume have been observed across the tree of life, with positive correlations documented between all three traits. Developmental tempo slows as genomes, nuclei, and cells increase in size, yet the driving mechanisms are poorly understood. To bridge this gap, we use a mathematical model of the somitogenesis clock to link slowed developmental tempo with changes in intra-cellular gene expression kinetics induced by increasing genome size and nuclear volume. We adapt a well-known somitogenesis clock model to two model amphibian species that vary 10-fold in genome size: Xenopus laevis (3.1 Gb) and Ambystoma mexicanum (32 Gb). Based on simulations and backed by analytical derivations, we identify parameter changes originating from increased genome and nuclear size that slow gene expression kinetics. We simulate biological scenarios for which these parameter changes mathematically recapitulate slowed gene expression in A. mexicanum relative to X. laevis, and we consider scenarios for which additional alterations in gene product stability and chromatin packing are necessary. Results suggest that slowed degradation rates as well as changes induced by increasing nuclear volume and intron length, which remain relatively unexplored, are significant drivers of slowed developmental tempo.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Society for Integrative and Comparative Biology.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Characterization of immunoglobulin loci in the gigantic genome of Ambystoma mexicanum.Front Immunol. 2023 Jan 27;14:1039274. doi: 10.3389/fimmu.2023.1039274. eCollection 2023. Front Immunol. 2023. PMID: 36776846 Free PMC article.
-
The kinetics in mathematical models on segmentation clock genes in zebrafish.J Math Biol. 2018 Jan;76(1-2):97-150. doi: 10.1007/s00285-017-1138-1. Epub 2017 May 25. J Math Biol. 2018. PMID: 28547212
-
Origin of amphibian and avian chromosomes by fission, fusion, and retention of ancestral chromosomes.Genome Res. 2011 Aug;21(8):1306-12. doi: 10.1101/gr.116491.110. Epub 2011 Apr 11. Genome Res. 2011. PMID: 21482624 Free PMC article.
-
A multi-cell, multi-scale model of vertebrate segmentation and somite formation.PLoS Comput Biol. 2011 Oct;7(10):e1002155. doi: 10.1371/journal.pcbi.1002155. Epub 2011 Oct 6. PLoS Comput Biol. 2011. PMID: 21998560 Free PMC article. Review.
-
Economy, speed and size matter: evolutionary forces driving nuclear genome miniaturization and expansion.Ann Bot. 2005 Jan;95(1):147-75. doi: 10.1093/aob/mci010. Ann Bot. 2005. PMID: 15596464 Free PMC article. Review.
Cited by
-
DNA gains and losses in gigantic genomes do not track differences in transposable element-host silencing interactions.Commun Biol. 2025 May 6;8(1):704. doi: 10.1038/s42003-025-08127-3. Commun Biol. 2025. PMID: 40328975 Free PMC article.
References
-
- Armstrong JB, Graveson AC. 1988. Progressive patterning precedes somite segmentation in the mexican axolotl Ambystoma mexicanum. Dev Biol 126:1–6. - PubMed
-
- Banfi S, Monti L, Acquati F, Tettamanti G, De Eguileor M, Grimaldi A. 2012. Muscle development and differentiation in the urodele Ambystoma mexicanum. Dev Growth Differ 54:489–502. - PubMed
Associated data
LinkOut - more resources
Full Text Sources
Miscellaneous
