Identification of ceRNA networks in type H and L vascular endothelial cells through integrated bioinformatics methods
- PMID: 39019785
- PMCID: PMC11255190
- DOI: 10.11817/j.issn.1672-7347.2024.230343
Identification of ceRNA networks in type H and L vascular endothelial cells through integrated bioinformatics methods
Abstract
Objectives: Type H blood vessels are a subtype of bone-specific microvessels (CD31hiEmcnhi) that play an important regulatory role in the coupling of angiogenesis and osteogenesis. Despite reports on the distinct roles of type H and L vessels under physiological and pathological bone conditions, their genetic differences remain to be elucidated. This study aims to construct a competitive endogenous RNA (ceRNA) network of key gene for differencial expression (DE) in type H and L vascular endothelial cells (ECs) through integrated bioinformatic methods.
Methods: We downloaded relevant raw data from the ArrayExpress and the Gene Expression Omnibus (GEO) database and used the Limma R-Bioconductor package to screen for DE lncRNAs, DE miRNAs, and DE mRNAs between type H and L vascular ECs. A total ceRNA network was constructed based on their interactions, followed by refinement using protein-protein interaction (PPI) networks to select upregulated and downregulated key genes. Enrichment analysis was performed on these key genes. Random validation was conducted using flow cytometry and real-time RT-PCR.
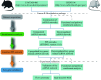
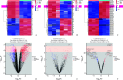
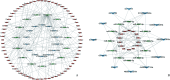
Results: A total of 1 761 DE mRNAs, 187 DE lncRNAs, and 159 DE miRNAs were identified, and a comprehensive ceRNA network was constructed based on their interactions. Six upregulated (Itga5, Kdr, Tjp1, Pecam1, Cdh5, and Ptk2) and 2 downregulated (Csf1r and Il10) key genes were selected via PPI network to construct a subnetwork of ceRNAs related to these key genes. Upregulated key genes were mainly enriched in negative regulation of angiogenesis and vascular apoptosis. Results from flow cytometry and real-time RT-PCR were consistent with bioinformatics analysis.
Conclusions: This study proposes a ceRNA network associated with upregulated and downregulated type H and L vascular ECs based on selected key genes, providing new insights into the regulatory mechanisms of type H and L vascular ECs in bone metabolism.
目的: H型血管是一种骨特异性微血管亚型(CD31hiEmcnhi),在成血管-成骨耦联机制中具有重要的调节作用。研究报道H和L型血管在骨骼生理和病理条件下发挥不同的作用,它们之间的遗传差异仍有待阐明。本研究旨在通过整合的生物信息学方法构建H和L型血管内皮细胞中关键差异表达(differential expression,DE)基因的竞争性内源RNA(competitive endogenous RNA,ceRNA)网络。方法: 从ArrayExpress和基因表达综合(Gene Expression Omnibus,GEO)数据库中下载相关原始数据,通过Limma R-Bioconductor包筛选H和L型血管间的DE lncRNAs、DE miRNAs和DE mRNAs,构建总ceRNA网络,随后根据蛋白质相互作用(protein-protein interaction,PPI)网络筛选出上调和下调的关键基因,以此精细化ceRNA网络并对关键基因进行富集分析。最后,通过流式细胞术和real time RT-PCR进行随机验证。结果: 共鉴定出1 761个DE mRNAs、187个DE lncRNAs和159个DE miRNAs,通过相互作用关系构建总ceRNA网络;通过PPI网络筛选出6个上调(Itga5、Kdr、Tjp1、Pecam1、Cdh5、Ptk2)和2个下调(Csf1r、Il10)的关键基因,并以此构建关键基因相关的ceRNA亚网络。上调的关键基因主要富集在血管生成与血管凋亡的负调控;流式细胞术和real-time RT-PCR的结果与生物信息学分析结果一致。结论: 本研究根据所筛选的关键基因提出上调和下调的H型和L型血管内皮细胞相关的ceRNA网络,为H型和L型血管内皮细胞在骨代谢中的调控机制提供了新见解。.
目的: H型血管是一种骨特异性微血管亚型(CD31 hiEmcn hi),在成血管-成骨耦联机制中具有重要的调节作用。研究报道H和L型血管在骨骼生理和病理条件下发挥不同的作用,它们之间的遗传差异仍有待阐明。本研究旨在通过整合的生物信息学方法构建H和L型血管内皮细胞中关键差异表达(differential expression,DE)基因的竞争性内源RNA(competitive endogenous RNA,ceRNA)网络。
方法: 从ArrayExpress和基因表达综合(Gene Expression Omnibus,GEO)数据库中下载相关原始数据,通过Limma R-Bioconductor包筛选H和L型血管间的DE lncRNAs、DE miRNAs和DE mRNAs,构建总ceRNA网络,随后根据蛋白质相互作用(protein-protein interaction,PPI)网络筛选出上调和下调的关键基因,以此精细化ceRNA网络并对关键基因进行富集分析。最后,通过流式细胞术和real time RT-PCR进行随机验证。
结果: 共鉴定出1 761个DE mRNAs、187个DE lncRNAs和159个DE miRNAs,通过相互作用关系构建总ceRNA网络;通过PPI网络筛选出6个上调( Itga5、 Kdr、 Tjp1、 Pecam1、 Cdh5、 Ptk2)和2个下调( Csf1r、 Il10)的关键基因,并以此构建关键基因相关的ceRNA亚网络。上调的关键基因主要富集在血管生成与血管凋亡的负调控;流式细胞术和real-time RT-PCR的结果与生物信息学分析结果一致。
结论: 本研究根据所筛选的关键基因提出上调和下调的H型和L型血管内皮细胞相关的ceRNA网络,为H型和L型血管内皮细胞在骨代谢中的调控机制提供了新见解。
Keywords: bioinformatics; bone vessels; ceRNA network; type H vascular endothelial cells.
Conflict of interest statement
The authors declare that they have no conflicts of interest to disclose.
Figures





Similar articles
-
Crosstalk Between H-Type Vascular Endothelial Cells and Macrophages: A Potential Regulator of Bone Homeostasis.J Inflamm Res. 2025 Feb 25;18:2743-2765. doi: 10.2147/JIR.S502604. eCollection 2025. J Inflamm Res. 2025. PMID: 40026304 Free PMC article. Review.
-
Identification of Key lncRNAs, circRNAs, and mRNAs in Osteoarthritis via Bioinformatics Analysis.Mol Biotechnol. 2024 Jul;66(7):1660-1672. doi: 10.1007/s12033-023-00790-3. Epub 2023 Jun 29. Mol Biotechnol. 2024. PMID: 37382793
-
Identification of Serum Exosome-Derived circRNA-miRNA-TF-mRNA Regulatory Network in Postmenopausal Osteoporosis Using Bioinformatics Analysis and Validation in Peripheral Blood-Derived Mononuclear Cells.Front Endocrinol (Lausanne). 2022 Jun 9;13:899503. doi: 10.3389/fendo.2022.899503. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 35757392 Free PMC article.
-
Analysis of ceRNA Network and Identification of Potential Treatment Target and Biomarkers of Endothelial Cell Injury in Sepsis.Genet Test Mol Biomarkers. 2024 Apr;28(4):133-143. doi: 10.1089/gtmb.2023.0143. Epub 2024 Mar 19. Genet Test Mol Biomarkers. 2024. PMID: 38501698
-
Molecular investigation of candidate genes for pyroptosis-induced inflammation in diabetic retinopathy.Front Endocrinol (Lausanne). 2022 Jul 25;13:918605. doi: 10.3389/fendo.2022.918605. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 35957838 Free PMC article.
Cited by
-
Crosstalk Between H-Type Vascular Endothelial Cells and Macrophages: A Potential Regulator of Bone Homeostasis.J Inflamm Res. 2025 Feb 25;18:2743-2765. doi: 10.2147/JIR.S502604. eCollection 2025. J Inflamm Res. 2025. PMID: 40026304 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

