Complete genome sequence of Microbacterium foliorum phage Curie, a podovirus isolated from soil in Spokane, Washington
- PMID: 39037314
- PMCID: PMC11320940
- DOI: 10.1128/mra.00408-24
Complete genome sequence of Microbacterium foliorum phage Curie, a podovirus isolated from soil in Spokane, Washington
Abstract
Bacteriophage Curie is a podovirus that infects Microbacterium foliorum. The Curie genome spans 16,810 bp, has 90 bp terminal inverted repeats, and includes 23 protein-coding genes. Its genome architecture resembles phage PineapplePizza and other phi29-like phages. Together, Curie and PineapplePizza form a new actinobacteriophage Cluster GI.
Keywords: bacteriophages.
Conflict of interest statement
The authors declare no conflict of interest
Figures
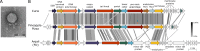
Similar articles
-
Complete annotated genome sequence of Microbacterium paraoxydans phage Evcara, a cluster GI podovirus isolated from compost.MicroPubl Biol. 2024 Dec 8;2024:10.17912/micropub.biology.001398. doi: 10.17912/micropub.biology.001398. eCollection 2024. MicroPubl Biol. 2024. PMID: 39717144 Free PMC article.
-
Complete genome sequence of phi29-like Microbacterium foliorum podovirus phage PineapplePizza.Microbiol Resour Announc. 2023 Oct 19;12(10):e0047823. doi: 10.1128/MRA.00478-23. Epub 2023 Sep 6. Microbiol Resour Announc. 2023. PMID: 37671874 Free PMC article.
-
Genome sequence of Microbacterium foliorum bacteriophage DumpQuist isolated from soil in Clarksville, Tennessee.Microbiol Resour Announc. 2023 Dec 14;12(12):e0092323. doi: 10.1128/MRA.00923-23. Epub 2023 Nov 22. Microbiol Resour Announc. 2023. PMID: 37991355 Free PMC article.
-
Complete Genome Sequence of Bacteriophage IndyLu, Isolated from a Microbacterium foliorum Culture.Microbiol Resour Announc. 2021 Dec 16;10(50):e0107921. doi: 10.1128/MRA.01079-21. Epub 2021 Dec 16. Microbiol Resour Announc. 2021. PMID: 34913713 Free PMC article.
-
Characterization and genome sequence of Microbacterium foliorum phage Morrigan, of cluster EA6 with siphovirus morphology.Microbiol Resour Announc. 2023 Dec 14;12(12):e0071923. doi: 10.1128/MRA.00719-23. Epub 2023 Nov 17. Microbiol Resour Announc. 2023. PMID: 37975671 Free PMC article.
Cited by
-
Complete annotated genome sequence of Microbacterium paraoxydans phage Evcara, a cluster GI podovirus isolated from compost.MicroPubl Biol. 2024 Dec 8;2024:10.17912/micropub.biology.001398. doi: 10.17912/micropub.biology.001398. eCollection 2024. MicroPubl Biol. 2024. PMID: 39717144 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
