Extracting fetal heart signals from Doppler using semi-supervised convolutional neural networks
- PMID: 39040082
- PMCID: PMC11260753
- DOI: 10.3389/fphys.2024.1293328
Extracting fetal heart signals from Doppler using semi-supervised convolutional neural networks
Abstract
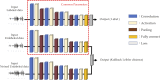
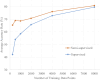
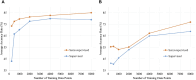
Cardiotocography (CTG) measurements are critical for assessing fetal wellbeing during monitoring, and accurate assessment requires well-traceable CTG signals. The current FHR calculation algorithm, based on autocorrelation to Doppler ultrasound (DUS) signals, often results in periods of loss owing to its inability to differentiate signals. We hypothesized that classifying DUS signals by type could be a solution and proposed that an artificial intelligence (AI)-based approach could be used for classification. However, limited studies have incorporated the use of AI for DUS signals because of the limited data availability. Therefore, this study focused on evaluating the effectiveness of semi-supervised learning in enhancing classification accuracy, even in limited datasets, for DUS signals. Data comprising fetal heartbeat, artifacts, and two other categories were created from non-stress tests and labor DUS signals. With labeled and unlabeled data totaling 9,600 and 48,000 data points, respectively, the semi-supervised learning model consistently outperformed the supervised learning model, achieving an average classification accuracy of 80.9%. The preliminary findings indicate that applying semi-supervised learning to the development of AI models using DUS signals can achieve high generalization accuracy and reduce the effort. This approach may enhance the quality of fetal monitoring.
Keywords: Doppler; deep learning; fetal heart signal; semi-supervised learning; ultrasound.
Copyright © 2024 Hirono, Kai, Yoshida, Sato, Kodama, Uchida and Kasai.
Conflict of interest statement
Authors YH and FU were employed by TOITU Co Ltd. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- Arnold J. J., Gawrys B. L. (2020). Intrapartum fetal monitoring. Am. Fam. Phys. 102, 158–167. - PubMed
-
- Chen L., Ruan W., Liu X., Lu J. (2020). “Seqvat: Virtual adversarial training for semi-supervised sequence labeling,” in Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics, 8801–8811.
LinkOut - more resources
Full Text Sources
Other Literature Sources

